@marpet
@ABraun
I would like to know how accurate is the LAI module in SNAP.
How does the accuracy of the product vary from deciduous to coniferous forest type?
I think this is hard to answer.
At first, it is not only the mathematical derivation but also the quality of the data and its radiometric calibration, but also the type of vegetation (a coniferous forest in Canada differs from coniferous forests in Portugal). So I would be very careful to draw fast conclusions on the quality.
I see this module similar to a supervised classification, for example. Technically, you can apply it to any data and class typology but in the end you are the only one who can tell if it makes sense because you know your study area best. And as for each accuracy assessment, you need field measurements for comparison.
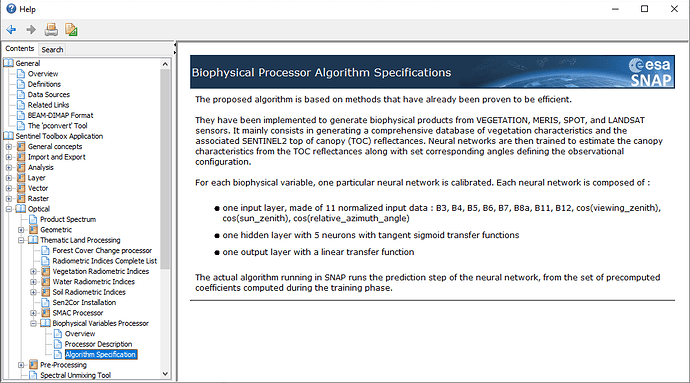
But you are right, the SNAP help does not tell much about the actual methods and their calibration. Maybe there is a document with a quality test around.
I could find these publications which tested the quality of the SNAP processor:
- Validation of the Sentinel Simplified Level 2 Product Prototype Processor (SL2P) for mapping cropland biophysical variables using Sentinel-2/MSI and Landsat-8/OLI data
- Comparison of SNAP-Derived Sentinel-2A L2A Product to ESA Product over Europe (4.4. Comparison of Vegetation Indices and Biophysical Parameters)
- Retrieval of crop biophysical parameters from Sentinel-2 remote sensing imagery
- Retrieval of Evapotranspiration from Sentinel-2: Comparison of Vegetation Indices, Semi-Empirical Models and SNAP Biophysical Processor Approach
- Canopy chlorophyll content and LAI estimation from Sentine1-2: vegetation indices and Sentine1-2 Leve1-2A automatic products comparison