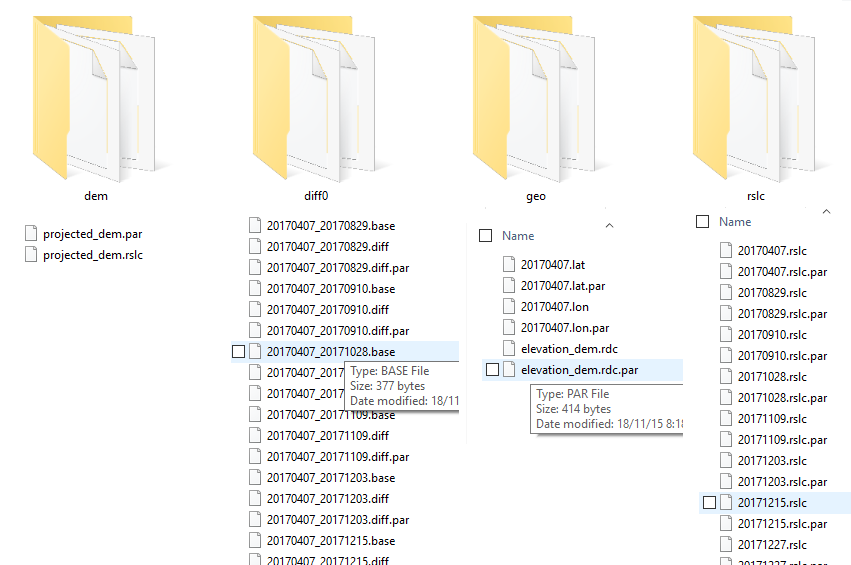
i forget it ,because the inf process stay almost 6 hours,and the error occurs,how can you process data (over 20 imgas),do i need two creat two more stack?
As I said, I do not know if it works if you split the data into separate stacks and I haven’t tested it.
Maybe you have access to a computer with more processing power to run it there.
Besides that, I can only recommend the snap2stamps tool again, which does all the data preparation for you after you modified the config files. Snap2stamps package: a free tool to automate the SNAP-StaMPS Workflow
thanks a lot,i am trying to create two stacks to process,and i dont know how to use the tools you have shared to me
these are simple python scripts. You need to have python installed and write in the config file where your master and slave data are located and then call the scripts.
A user manual is contained in the download.
The end is the prepared dataset which is then used for PS processing.
thanks a lot but i found it is a little difficult
the great advantage is that you can go sure that the result is ready for PS InSAR processing while working with several stacks could still lead to some errors at a later step.
Plus, it processes the files separately, so it might also be more efficient regarding your system’s capacities.
ok i will try it
it seems in the manual of the tools ,i need to use linux OS instead of windows,am I ?
no, it works under Windows as well. You only have to switch to Linux afterwards. If you have questions on this tool, please use the mentioned topic to keep the discussions clean.
Hi all @thho @ABraun @Fikretjfm
Somebody know why I get this message when I want run the script mt prep gamma???!!!
200 does not exist, exiting
Thanks for advance
try to rename the folder “processed data” to “processed_data” and run again. It could be that the script expects the variables in the exact number and order and the whitespace caused the error.
good job!
Getting into StaMPS and its preprocessing takes its time, but the possibilities and results are quite rewarding.
I need new results!!! 


Another way to avoid this problem is to pass the path-string in quotation marks, or use \ before the white space to escape it, this can also be used in order to escape signs like $ which do have other meanings in shell scripts…anyway avoiding white space and special sings in file and directory names is always recommend.
HI
To calculation PSI in SNAP 6.0 I did below Steps:
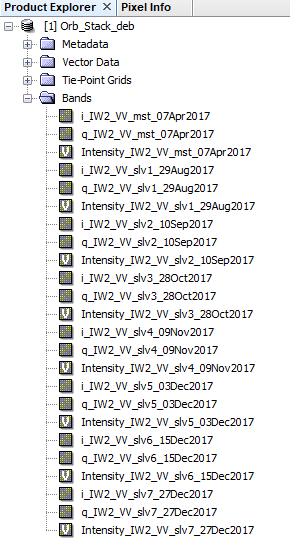
1- Prepared the Orb_Stack_deb
including:
2- prepared the Orb_Stack_deb _inf_dinsar
Including : elevation and orthorectifiedLat and orthorectifiedLon
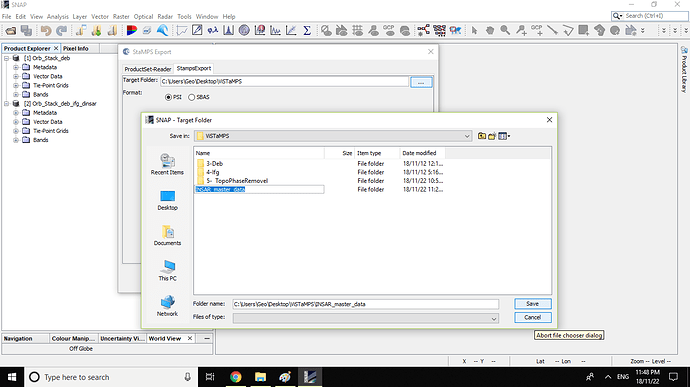
3- Export them by SNAP ‘‘StaMPS export tool’’ into “INSAR_master_data” folder
4- Now i have this folders and their contents
5- Please guide me to the next steps of STaMPS
My questions are as follows:
Command mt_prep_gamma (in StaMPS-4.1-beta\bin) In what software and how it runs?
Should I get this folder (13 gigabytes) into linux and run it there? How?
Should I run with Cygwin64 ? How?
The next step in the software should run? How?
Please provide detailed guidance
I read all post in topic but can not STaMPS !!!
Thanks in advance
I don’t recommend cygwin. If you have the chance to use Linux for the next steps, download the StaMPS scripts there and set the variables of the other software (you need matlab, triangle and snaphu, the rest is only used for pre-processing which was done in SNAP)
Once you have these programs installed, you source the config file as described in the StaMPS manual or here:
- Linux Installation using StaMPS and S-1 data
- About the STaMPS category
- Workflow between SNAP and StaMPS
- Problem with StaMPS installation using Linux
mdelgado also explained it well here: How to prepare Sentinel-1 images stack for PSI/SBAS in SNAP 5
If everything is installed, you call the scripts to prepare the data for matlab processing as described in the manual under chapter 6 PS processing.
Hi ABraun and thank U for response
unfortunately, i not know how run " mt_prep_gamma" and which software is need to ran it?
is it possible to you mention the steps after stages above(step by step).
Should I copy these folder (about 13 gigabytes) into linux ?
i copied these folder to linux but can not run " mt_prep_gamma"
thanks
No software, these are linux scripts. You open your shell and type in the commands.
Maybe someone at yout place who is experienced with Linux can assist you. I think this is beyond the scope of this forum, but it has been explained here several times.
Maybe also this is a start.
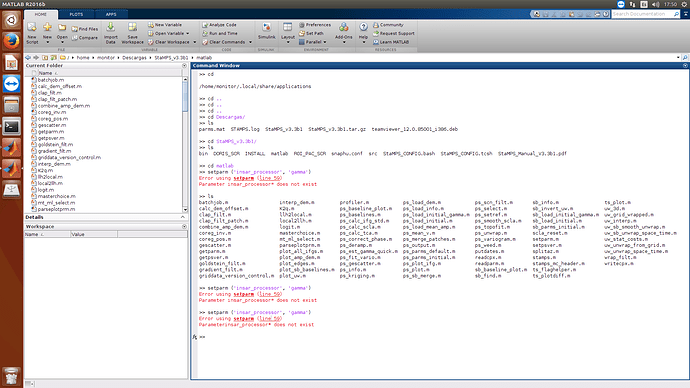
Hi all @ABraun @thho @Fikretjfm
In the end of my processing I have problem with MATLAB, please help! 


Parameter insar_processor* does not exist!!!
Thanks for advance