Does anyone knows how to subset an image in Snap without this problem?
It is a S2 scene, right?
In which data format have you saved the subset?
Yes it’s a S2 scene.
It’s not about format, it happens before i export the image to any format.
I want to export the subset to tiff format, but subset destroys the image.
My solution for now is to export the whole image to envi format, and i do subset in envi.
But this procedure need a lot of time and it’s not a good solution for me.
This could be a histogram-stretching issue. The subset has different statistics than the whole scene so the default stretching could be different?
I have no idea what’s happening. There are no “options” in subset window, so you can’t try different options.
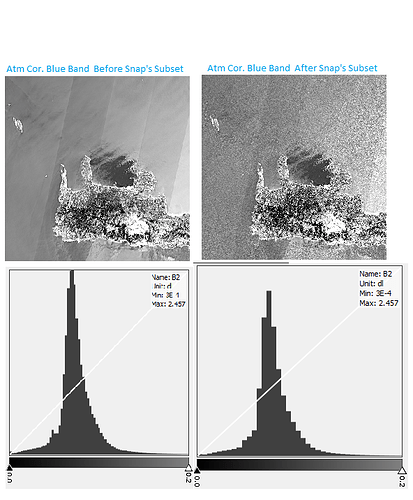
As you can see the subset area is exactly the same with the starter image, so there isn’t any logical reason to change the values of pixels. I just removed individual bands without to do any changes to spatial area, and it destroy the image…
I just did it and the values were the same as before the subset. But I had only a land scene at hand.
Can you set some pins and see how the values change.
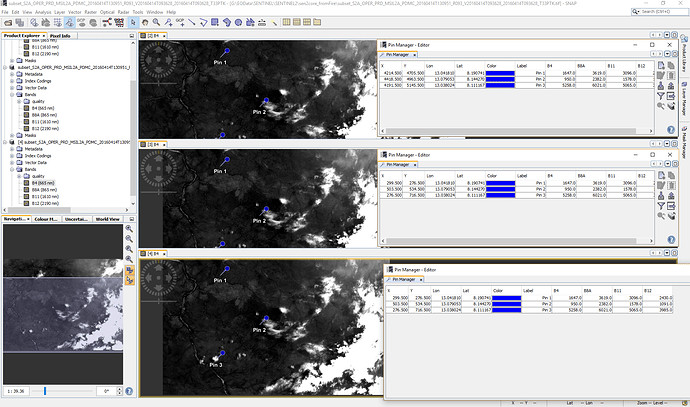
In the following image you see what was shown for me.
The first is the original, the second before saving to tif and the third is the reopend tif.
have you tried to open the data with the S2 reader (File > Import >
Optical > Sentinel-2 > Level 2A) and thus skip the manual resampling by
choosing an UTM zone and a resampling level?
Have you tried to zoom into the scene? Are the images equal when zooming in?
What I just did, is to resample one product and then create a band subset, just like you did.
Afterwards I also saw differences. So I subtracted both images from each other by Band Maths.
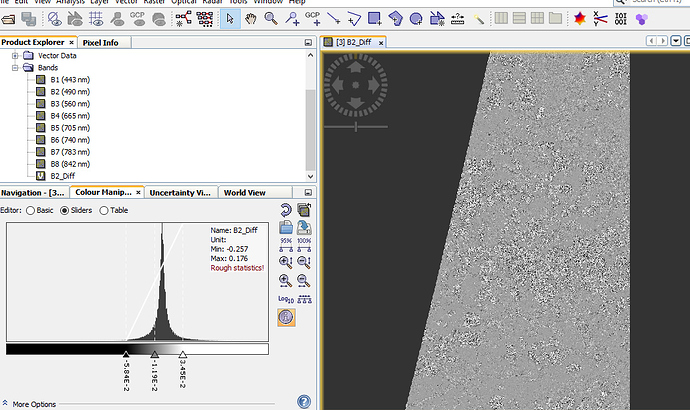
The result you can see here:
You see that the image is noisy and also the histogram shows differences. The image shows the image data of level 3 of the image pyramid.
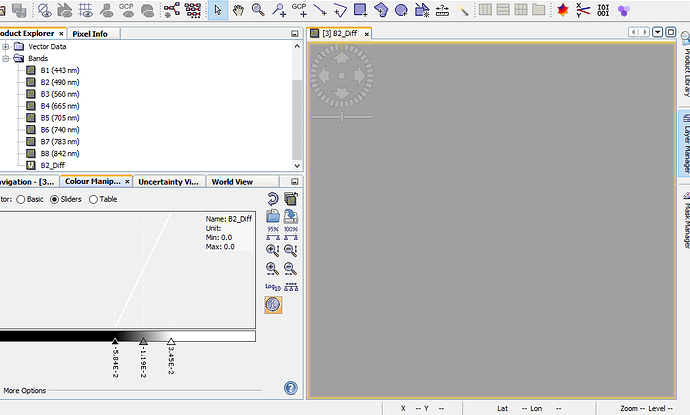
But when I zoom in to level zero, the images becomes smooth and if I calculate the exact statistics I see that all values are zero.
So the problem is more a problem of displaying the data as actually the data itself. So using the data and further processing it is save.
Nonetheless, this should be considered as a bug. I’ve tried it with MERIS and Landsat8. With MERIS I saw the same effect but not in Landsat8. I’m not sure where it comes from. Probably not only related to the subset operation but also to the data reader and the resampling operation. Maybe from different resampling strategies before and after the subset.
This should be further investigated. I’ve put it into our issue tracker.
Thanks a lot for your report.