This is what I did:
- Download two S2 products (same tile) and extract them
- Apply S2 Resampling to bring the product into a common resolution
- Create a subset for each (same area)*
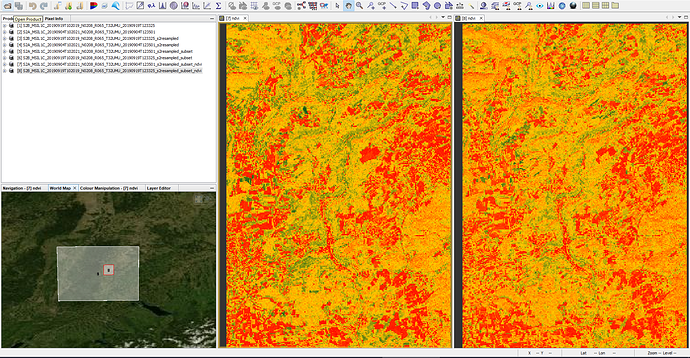
- Perfom the NDVI operator
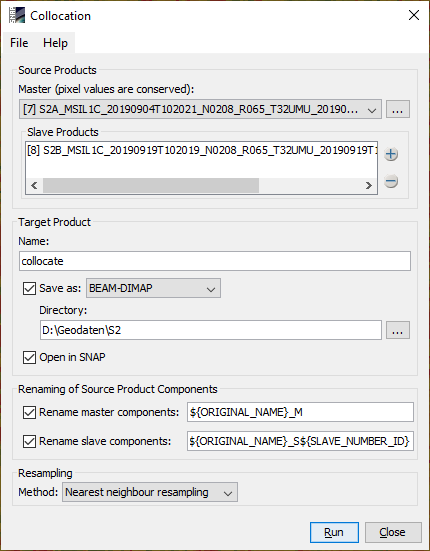
- Collocate both NDVI products (get the same error, I don’t understand it)
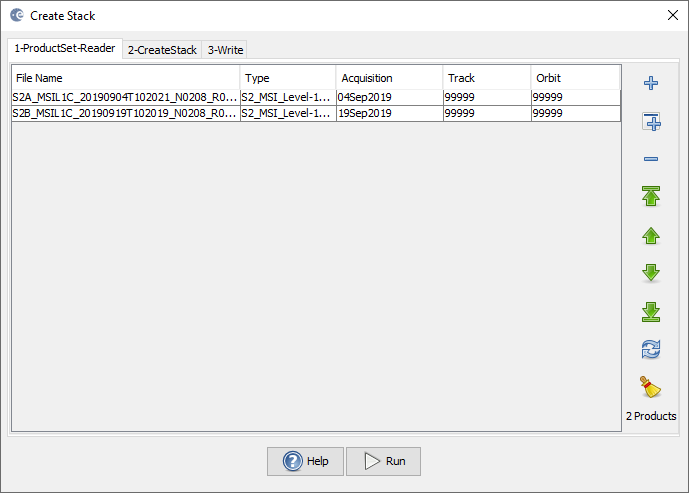
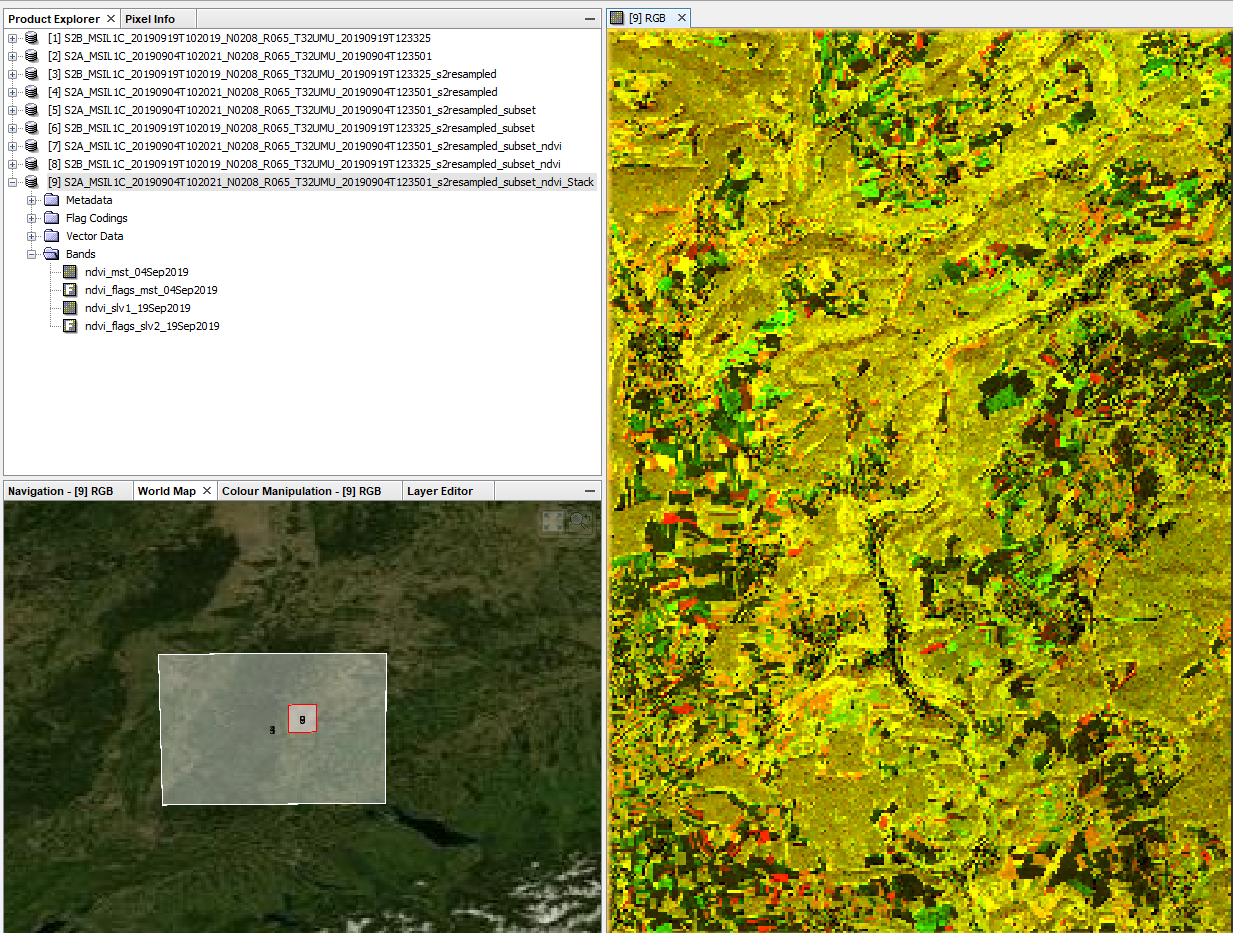
- Alternative: Use Create Stack (Radar Coregistration > Stack Tools), select Product Geolocation instead of Orbit in the second tab
*whenever you create a subset you have to make sure that it is correctly written and that all bands are correctly transferred into the subsequent product. Sometimes, SNAP “loses” some of the bands during S2 subsetting.
NDVI images
Collocation
Collocation error message
Stack