Thanks for the advice.
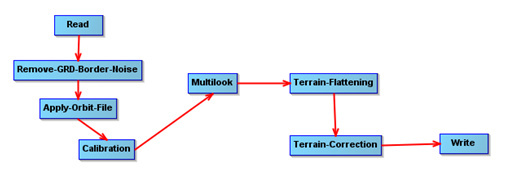
I want to apply speckle-filter to the subset image because it will be very slow for the whole tile, and for now we only need sampling based analysis. For example, we extract information from a set of 20x20m sampling blocks (75 blocks in total). Each block has a series of Sentinel-1 revisit (IW GRDH). My pre-processing chain is saved in the graph below:
VVVH_Rb_Orb_Cal_M2_TF_TC_SNAP20m.xml (5.8 KB)
Then I subset and stack back using gdalwarp and gdal_merge, which is seems more convenient for bash scripting compared to using SNAP via gpt.
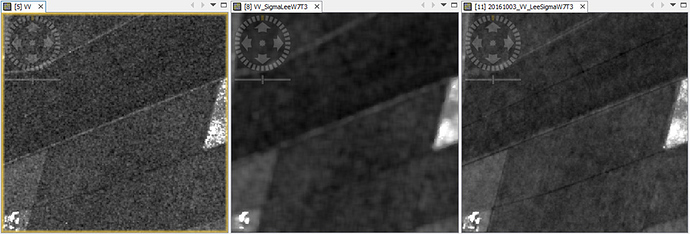
There’s no problem when applying single-product filtering (e.g. Lee Sigma). I would like to apply the multi-temporal Lee Sigma filter because this http://forum.step.esa.int/t/single-or-multi-temporal-speckle-filter/2139 suggest to use it. I also did a quick run over one of the sampling block, and the results prove that Multi-temporal Lee Sigma Filter performed the best (see image below).
(left: Raw image, Gamma0_VV, power scaled; mid: single-product Lee Sigma filter; right: multi-temporal Lee Sigma filter).
However, the multi-temporal filter required this “Unit” info which lost during the process of subseting (via gdalwarp) and stacking (via gdal_merge.py). Above Multi-temporal filter result was derived by manually add “Unit” back for each band in stack.
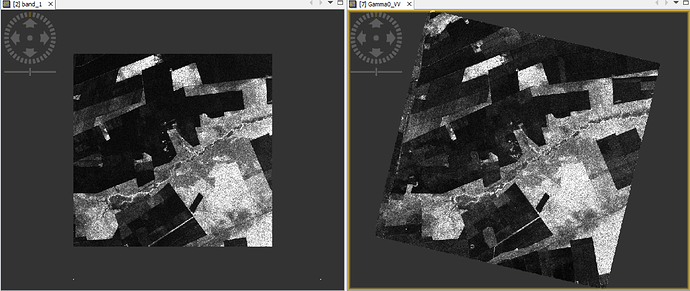
Also I test to do “Subset” after “Apply-Orbit-File and Calibration”, but before other pre-processing (e.g. “Multilook”, “Terrain-Flattening”, and “Terrain-Correction”). I see there is an option to use “Geographic Coordinates”, and the test under SNAP/GUI works. But it seems there are some issues that the subset is not in square (from geojson to WKT using http://geojson.io/):
In sum, my questions are:
A) Can we add the metadata back for subset image stack, say from the pre-processed Gamma0 image? It seems using WKT to subset then I will need to use gdalwarp again to get a square output in my case.
B) If must do the within SNAP, is there an option to give input vector (shapefile/geojson) for “subset”? As mentioned above, I will need to apply this in a batch mode, using"gpt" via bash script.
Thanks!