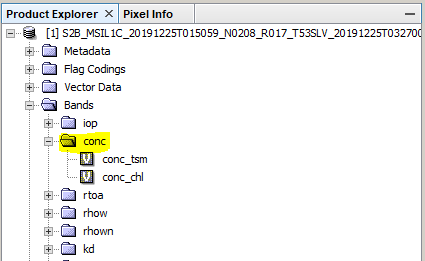
the chlorophyll and suspended matter concentrations are in the folder Bands > conc
I found conc_tsm and conc_ chl, but no image is displayed.
Is there not enough RAM?
you can check with the statistics operator if the band has values (then it is s problem of RAM) or not (then it was not computed successuflly)
I wonder why the band is not stored under “conc”
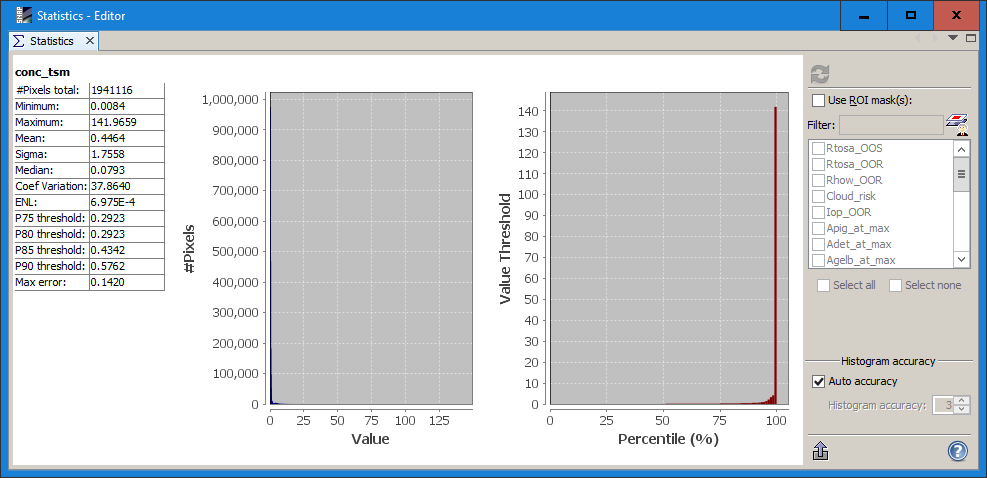
To see if there are values in your raster, you can use the statistics tool ![]()
It should show values in the table and the histogram:
while an empty raster will not have any data here.
Until now, any data has never been divided into files.
When I started the statistics tool, nothing was displayed.
There seems to be some processing that cannot be done with L2A. Is there any processing that cannot be done with L1C?
as I said, the C2RCC needs L1C as an input, not L2A.
You can apply any processing to L1C products (as long as they are resampled to a common resolution)
hi, i want to ask about C2RCC processor parameters how can i find ozone and air pressure at sea level values for Turkey?
There are several sources. You could try the following
Trying to run C2X-Complex-Nets from Snappy, but it keeps throwing "RuntimeError: org.esa.snap.core.gpf.OperatorException: Operator ‘C2rccMsiOperator’: Value for ‘Set of neuronal nets’ is invalid: ‘C2X-COMPLEX-Nets’. Any idea why?
C2X-Nets and C2RCC-Nets do work.
That’s strange. In the code this is the expected name.
Can you show your related code?
I know, right? I got the parameter from both the source code and gpt -h. Here is my code, the problem is at the end:
import sys
import os
sys.path.append('/home/sergio/.snap/snap-python')
from esa_snappy import ProductIO, GPF, HashMap, WKTReader
# Directorio con archivos de entrada y de salida
input_directory = '/home/sergio/PycharmProjects/tfg/downloaded_SAFE'
output_directory = '/home/sergio/PycharmProjects/tfg/ProcesadosC2X-COMPLEX'
# El directorio existe
os.makedirs(output_directory, exist_ok=True)
# Coordenadas de la región de interés
wkt = 'POLYGON ((-0.915298 37.610152, -0.915298 37.835819, -0.671883 37.835819, -0.671883 37.610152, -0.915298 37.610152))'
geom = WKTReader().read(wkt)
# Obtener lista de archivos en el directorio de entrada
for file_name in os.listdir(input_directory):
if file_name.endswith('.SAFE'):
input_file = os.path.join(input_directory, file_name)
output_file = os.path.join(output_directory, f'processed_{file_name}')
print(f'Procesando: {input_file}')
# Leer el producto de entrada
source_product = ProductIO.readProduct(input_file)
# Resampling
parameters = HashMap()
parameters.put('targetResolution', '20') #TODO: si lo bajo a 10 pasa a tardar 1h/producto, pero podría mejorar
parameters.put('geoRegion', geom)
resampled_product = GPF.createProduct('Resample', parameters, source_product)
# Recorte (que en principio ya está hecho, pero no sé si puede aplicar el resample a la vez que el ajuste de la zona)
parameters = HashMap()
parameters.put('geoRegion', geom)
subset_product = GPF.createProduct('Subset', parameters, resampled_product)
# Parámetros C2RCC
parameters = HashMap()
parameters.put('salinity', '35.0')
parameters.put('temperature', '15.0')
parameters.put('ozone', '330')
parameters.put('press', '1000')
parameters.put('elevation', '0')
parameters.put('TSMfac', '1.06')
parameters.put('TSMexp', '0.942')
parameters.put('CHLexp', '1.04')
parameters.put('CHLfac', '21.0')
parameters.put('outputAsRrs', 'false')
parameters.put('outputRtoa', 'true')
parameters.put('outputAcReflectance', 'true')
parameters.put('outputRhown', 'true')
parameters.put('outputKd', 'true')
parameters.put('outputUncertainties', 'true')
parameters.put('netSet', 'C2X-COMPLEX-Nets') # Cambiando esto se aplican distintas redes neuronales: C2X-Nets, C2RCC-Nets
# Aplicar el procesador C2RCC
c2rcc_product = GPF.createProduct('c2rcc.msi', parameters, subset_product)
# Guardar el producto
ProductIO.writeProduct(c2rcc_product, output_file, 'BEAM-DIMAP')
print(f'Archivo procesado guardado en: {output_file}\n')
print('Procesamiento completado.')```From the GUI the parameter works. I currently have no snappy setup. So, I can’t reproduce this.
Have you tried str('C2X-COMPLEX-Nets')?
It should work without it, but who knows?
Didn’t work, it outputs the same error. Do you know anyone else with a snappy setup that could try and reproduce the error? I really need to get this to work.
Maybe @dolaf or @TomBlock can try.
If as also made a pull request to enhance the error message.
If accepted, the error message will include the accepted values of the value set, like “Value for ‘X’ is invalid: ‘d’. Allowed values are: [a, b, c].”
It seems the devs do not have the time or don’t want to reply.
I have now set up a snappy environment for SNAP 12. I’ve used a miniconda environmment and Python 3.12.9.
The script worked. I’ve only changed the region and removed the geom from the resampling as it is not used by this operator.
Also, you should use the S2Resampling. It improves the results of the C2RCC. And if you not really need the uncertainties, disable them. They are computational intense.
Unfortunately, I can’t reproduce your error.
import os
from esa_snappy import ProductIO, GPF, HashMap, WKTReader
# Directorio con archivos de entrada y de salida
input_directory = 'D:\\EOData\\_temp\\c2rcc'
output_directory = 'D:\\EOData\\_temp\\c2rcc\\output'
# El directorio existe
os.makedirs(output_directory, exist_ok=True)
# Coordenadas de la región de interés
# wkt = 'POLYGON ((-0.915298 37.610152, -0.915298 37.835819, -0.671883 37.835819, -0.671883 37.610152, -0.915298 37.610152))'
wkt = 'POLYGON ((-127.46064834768475 49.835330370524886, -127.15343986701647 49.830842294687784,-127.16162293324992 49.614676446975885, -127.46747144562684 49.61913054266424,-127.46064834768475 49.835330370524886))'
geom = WKTReader().read(wkt)
# Obtener lista de archivos en el directorio de entrada
for file_name in os.listdir(input_directory):
if file_name.endswith('.SAFE'):
input_file = os.path.join(input_directory, file_name)
output_file = os.path.join(output_directory, f'processed_{file_name}')
print(f'Procesando: {input_file}')
# Leer el producto de entrada
source_product = ProductIO.readProduct(input_file)
# Resampling
parameters = HashMap()
parameters.put('targetResolution', '20') # TODO: si lo bajo a 10 pasa a tardar 1h/producto, pero podría mejorar
# parameters.put('geoRegion', geom) # Does not have this parameter
# better use S2Resampling - improves C2RCC results
resampled_product = GPF.createProduct('Resample', parameters, source_product)
# Recorte (que en principio ya está hecho, pero no sé si puede aplicar el resample a la vez que el ajuste de la zona)
parameters = HashMap()
# parameters.put('geoRegion', geom)
subset_product = GPF.createProduct('Subset', parameters, resampled_product)
# Parámetros C2RCC
parameters = HashMap()
parameters.put('salinity', '35.0')
parameters.put('temperature', '15.0')
parameters.put('ozone', '330')
parameters.put('press', '1000')
parameters.put('elevation', '0')
parameters.put('TSMfac', '1.06')
parameters.put('TSMexp', '0.942')
parameters.put('CHLexp', '1.04')
parameters.put('CHLfac', '21.0')
parameters.put('outputAsRrs', 'false')
parameters.put('outputRtoa', 'true')
parameters.put('outputAcReflectance', 'true')
parameters.put('outputRhown', 'true')
parameters.put('outputKd', 'true')
parameters.put('outputUncertainties', 'true')
parameters.put('netSet',
'C2X-COMPLEX-Nets') # Cambiando esto se aplican distintas redes neuronales: C2X-Nets, C2RCC-Nets
# Aplicar el procesador C2RCC
c2rcc_product = GPF.createProduct('c2rcc.msi', parameters, subset_product)
# Guardar el producto
ProductIO.writeProduct(c2rcc_product, output_file, 'BEAM-DIMAP')
print(f'Archivo procesado guardado en: {output_file}\n')
print('Procesamiento completado.')
Dear Marco, thank you very much for the advice.
I just solved the problem by reinstalling the whole Sentinel 3 toolbox. I believe the error comes from installing an older version of the project. By following the instructions in s3tbx-c2rcc, I got misleaded by the “How to build” section, which holds a link to the deprecated C2RCC processor github page. This version of the processor happens to be older than the release of C2X-COMPLEX nets, thus not recognizing the appropriate parameter.
Again, thank you for your efforts and I apologize for making you lose time over this.
No worries. Great that it now works for you.
Thank you, i also wonder, is C2RCC working with S2-L2A (BOA)