I have resampled the image to 20 m, but if I apply idepix tool to masking cloud. it gives me the following error:
do you have solution about that? thanks before!
I have resampled the image to 20 m, but if I apply idepix tool to masking cloud. it gives me the following error:

do you have solution about that? thanks before!
Some time ago this has been reported for a different use case.
Please have a look at this issue ticket (SNAP-561). Especially at the comment below.
Good night Sir,
Is the similar procedure for Sentinel 2A Level-2A?
Atmospheric correction applied to Top-Of-Atmosphere (TOA) Level-1C. So, the Level-2A output is an orthoimage Bottom-Of-Atmosphere (BOA) corrected reflectance product(https://sentinel.esa.int/web/sentinel/user-guides/sentinel-2-msi/processing-levels/level-2). The IdePex Sentinel 2 MSI have an option for write ToA reflectance to the target product and other option for write feature values to the target product. What to do in this case?
Thank you for your valuable response.
IdePix doesn’t work on L2A data. You can run it on L1C and then merge the result with the L2A data.
Or you can use the cloud mask of the L2A product directly.
Thank you Sir,
Do you have a link of a tutorial for cloud mask the L2A using the second option? Grateful for your advise.
You can use the masks provided with the data.
For example in the valid-pixel expression:
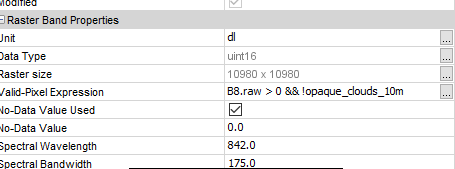
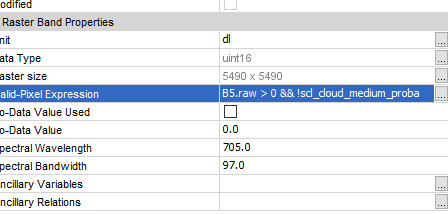
Maybe this is also of interest for you:
Sir I do not understand. I want to do this with SNAP. I did it before in Google Earth Engine with level 1C. I am just no familiar with this software. I used the option in gis.stackexchange.com as you advised using a virtual band I created in the mask manager. I got this. What is my next step to get the bands than I need free of cloud cover?
Please.
If you have the cloud mask it depends on your final goal what will be the next step.
For an RGB image with out clouds you can modify the valid expression for each of the used bands.
For example:
B2.raw > 0 && !scl_cloud_medium_proba && !scl_cloud_high_proba
And the same for B3 and B4.
Image before:
Image after:
Thank you Sir,
My goal is to computer 4 spectral indexes:
The Modification of the Normalised Difference Vegetation Index
The Normalised Difference Vegetation Index (NDVI)
The Normalised Difference Bareness Index (NDBal) and
The Normalised Difference Built-up Index (NDBI).
At the end I have to merge all the layers together and compute the unsupervised classification K-Mean for a bauxite mine site. So I need the the SWIR band Band 11 to be resampling to 10m resolution I also need a VNIR band, in this case I can used Band 8.
Grateful for your advise,
Susy
Then do the modification for band B11 and B8.
B8.raw > 0 && !scl_cloud_medium_proba && !scl_cloud_high_proba
B11.raw > 0 && !scl_cloud_medium_proba && !scl_cloud_high_proba
Or do it in the NDVI expression when using the Band Maths:
if (scl_cloud_medium_proba || scl_cloud_high_proba) then NaN else ((B8 - B11) / (B8 + B11))
Thank you Sir. You are my only help in this entire country.
Good night Mr Marpet,
I did the cloud masking for each bands. I have to do something else with the image in the future.
Please find attached the results. I want to see the free cloud cover image in QGIS or SAGA GIS as I used to do with GEE images. When I load the raster image into QGIS after the cloud masking. The clouds still there?
Yes, when exporting the bands with such expressions it gets probably lost. Other applications don’t know it.
But you can create new bands which you can then export.
For the RGB channels at the example of B4 (red):
red = if scl_cloud_medium_proba || scl_cloud_high_proba then NaN else B4
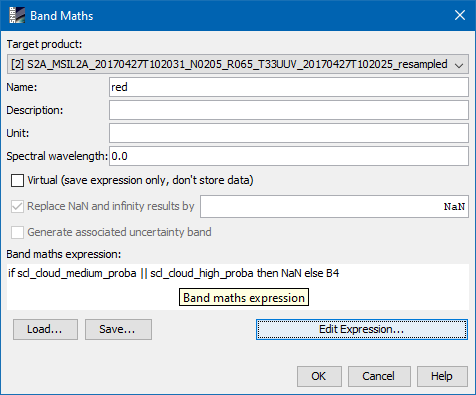
Make sure to disable the virtual option.
Good morning Mr Marpet,
Thank you for all your help. Do I need more than one product to create a mosaic of images?
I am thinking of what do to after removing the clouds?
Do I need to reconstruct the spaces remaining?
Resampling the whole image area take a lot of time, Is it possible to created a subset of the area of interest prior to resampling the sentinel product?
I am using one single product for a specif time.
Regards
Yes, for a mosaic you need more then one product.
You can use multiple from the same sensor for the area but from different dates.
But if you are interested in a specific date you have to go one with the clouds, respectively with the wholes.
There are also techniques to merge data from multiple sensors. But this is out of my scope. The data needs to be pre-processed properly.
Yes, you can first crop the scene to the area you are interested in and then do the resampling.
Maybe the S2GM service can help you. it provides Sentinel-2 L2A mosaics globally.
https://land.copernicus.eu/imagery-in-situ/global-image-mosaics/node/16
Hi forum I have an NDVI image from sentinel 2 so the value ranges from -0.9 to +0.9 but I want an NDVI value which ranges from 0.1 to 0.8 so I made a mask using mask manager however I don’t how I can change the masked one to the band form. Additionally please I get confused about the two data types of sentinel 2 which are S2 MSI1C and S2 MSI 2A. does it mean we can directly use S2 MSI 2A without Atmospheric correction and cloud cover reduction? Again for S2 MSI1C after changed into S2 MSI 2A using the Sen2 core processor can we use it for further analysis? Thank you all.
You can now open the band properties of the NDVI and enter the name of the mask in the “Valid pixel expression”. This will remove all non-mask pixels from the NDVI raster. It is dynamic, so whenever you change the mask, the NDVI will change accordingly.
For the other question: L1C is top-of-atmosphere (TOA) reflectance, L2A is bottom of atmosphere (BOA) reflectance.
So if you apply atmospheric correction to L1C you get L2A.
Thank you ABuran so do you mean if I have L2A I don’t need to make corrections
yes. It’s not perfect, but the data is atmosperically corrected for sun angle, shades and slight haze.
HI, Abraun I have another question I have 2 SAR imagery and 2 Sentinel 2 NDVI images. I have masked out unnecessary pixels individually for each image but I want to mask out again the 4 images at the same time I mean if there is NaN value pixel in the NDVI I want it to be in the corresponding SAR imagery too