Hello,
I am using Sentinel-3 SYN L2 AOD product. I want to covert some parameters of the product to Geotiff format.
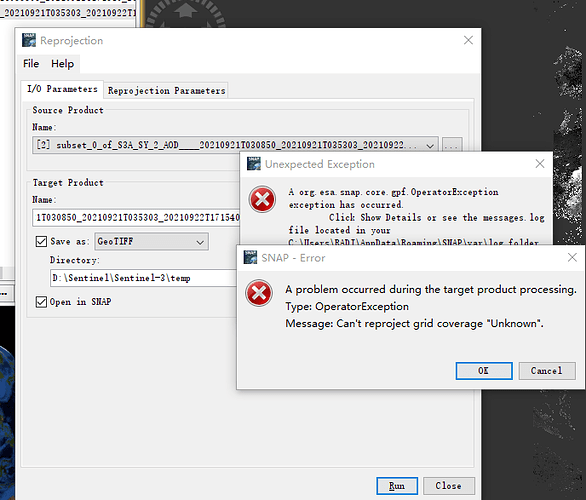
Now, firstly, I open the xfdumanifest.xml in snap. Then, I use Raster/Subset to extract some parameters(AOD550nm, lon, lat). Finally, I use reprojection operation (Raster/Geometric Operations/Reprojection) to reproject and export this to Geotiff (WGS-84). But now, I I encountered this error: (Can’t reproject grid coverage “Unknown”.
That’s strange. I haven’t seen it before.
The error message seems to come from a library we use.
Can you provide the log file?
The link explains how to find the logs: SNAP FAQs - How to find the log files?
I had the same problem, my solution was: subset(all the data), reproject and extract bands and save as tiff.
I don’t remember if I found other solution. 
Hi,
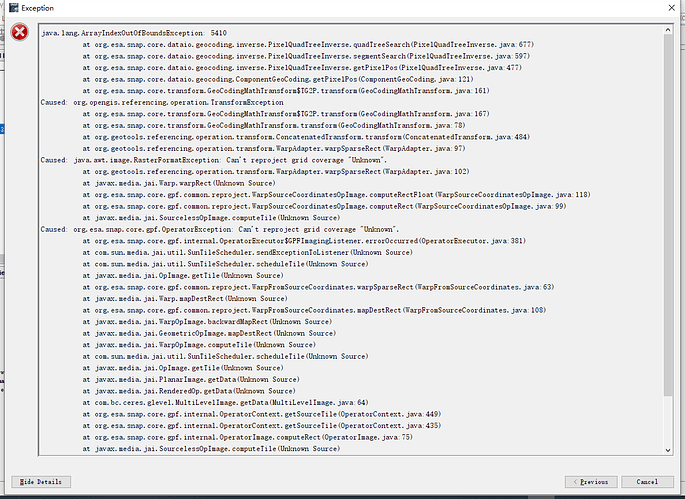
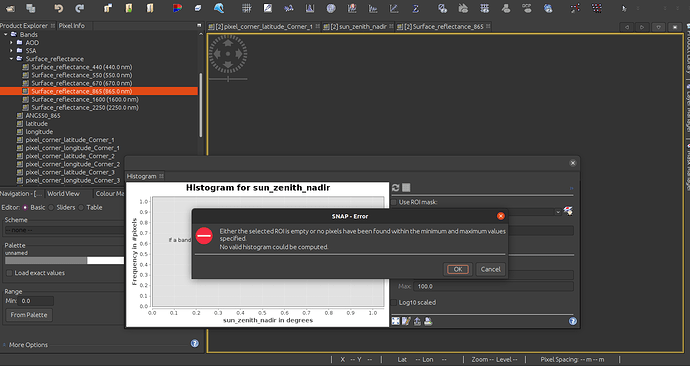
Thanks for taking the time to help with questions! Firstly, this is the detail of this error:
This is the log file:
messages.log (80.9 KB)
In addition, I want to know if there are other ways to convert this product to Geotiff format?
Yours sincerely,
Hao Zhu
Hi,
Thanks for your reply! It does not work on me. As long as I execute subset, I will encounter this error, even if I select all data when executing subset.
Maybe you can try save on netcedf and after export as tiff. You try reproject before subset?
Thanks for posting the exception. This helped to identify the reason.
This happens due to a bug in in the current release. It is already fixed for the next major release.
It would be great if you can workaround this issue by changing the configuration of the reader.
Please add the following line to the s3tbx.properties file.
s3tbx.reader.syn.aod.pixelGeoCoding.inverse=INV_PIXEL_GEO_INDEX
The file can be found in .snap/etc in the user directory. If it does not exist yet you can create it.
With this setting the error should not happen anymore.
Thanks for your reply! I added this line to the s3tbx.properties file. The error disappeared!But the result was still incorrect.
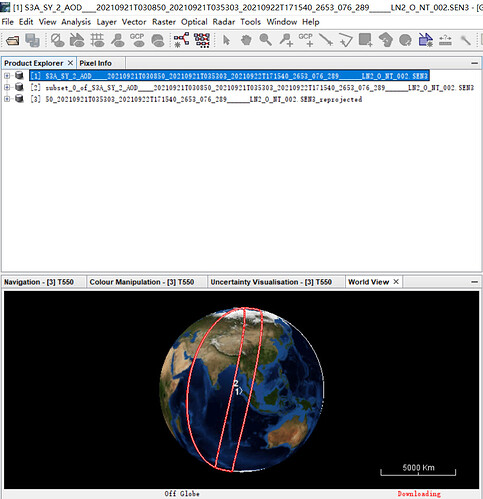
This is original data:
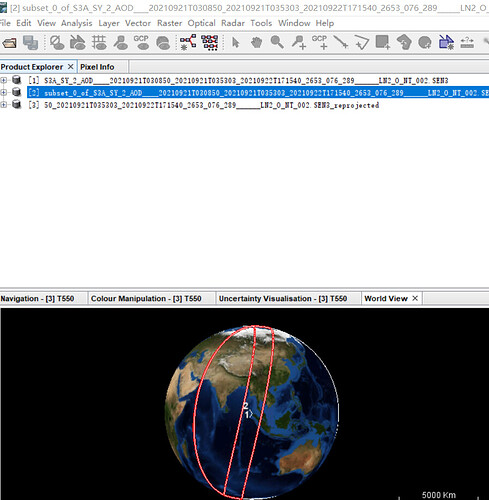
This is the subset data:
This is the reprojected data:
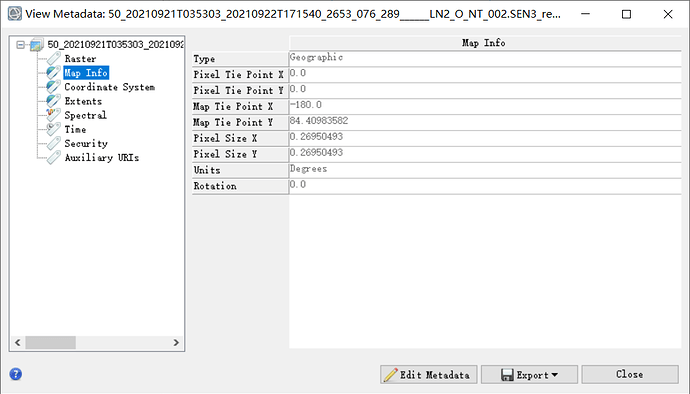
Obviously, the display range of the results is incorrect. I opened it by ENVI:
The resolution is also incorrect: the true resolution is 4.4km
Hi, i have a problem also working with Sentinel-3 SYN L2 AOD. When i try to reproject (Raster > Geometric > Reprojection )the layer it returns a NaN dataset. I attached the logs. I did the process in SNAP 9.0.0 on linux (Ubuntu 20.04) with 16 GB ram
messages.log (78.3 KB).
Anyone knows what could it be happening?
Best regards,
Italo
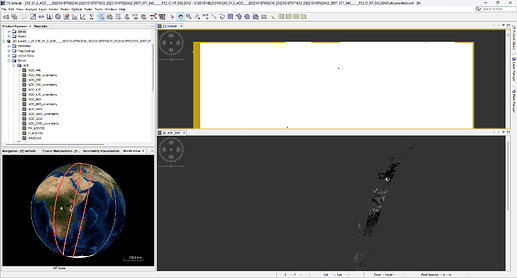
Hi this is cause due to no-data (NaN) values in the latitude and longitude data.
The no-data is at the borders. The yellow area in the below.
These no-data values cause also the strange boundary on the globe.
It can be fixed if you create a subset first.
In my case I excluded ~15 left and right and 2 pixels at the top. I don’t know if this constant for all products. With a bit more margin this can automatically applied in an operational processing chain. Afterwards the reprojection worked.
Thanks for your response and support Marco. Effectively now it works ![]() !
!