Hi, my name is Alex and I’m new to Snap.
I’m currently in the development of my diploma license on processing Sentinel 2 L1C images to derive chlorophyll concentration.
I have managed to process the images to 10 m, make subset of them and apply c2rcc processor. I am happy with the result.
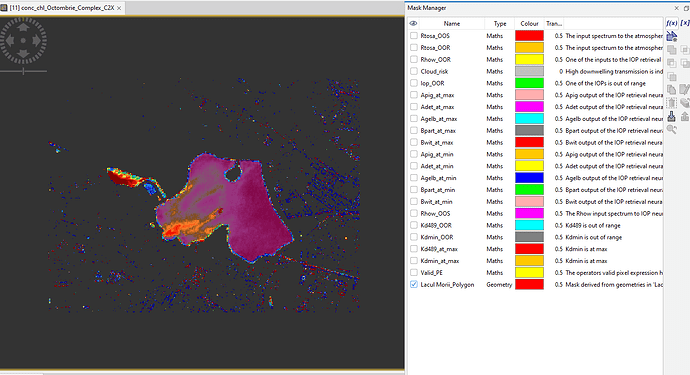
However some random pixels have not been masked and I am having trouble finding ways to delete the unwanted pixels.
I have created a polygon for the area of interest and I am trying to find out how to use the mask option to export only the selected area.
Can you please give me an advice on how to proceed?
My final goal is to export the area into Google Earth KMZ, only the area, not the entire view.
Hi Alex,
you can use the mask directly in the valid-pixel expression of the conc_chl band.
Right-click the band → select Properties and set the expression to c2rcc_flags.Valid_PE && ´Lacul Morii_Polygon´
Better don’t use Space in the Mask name. You have to escape it. It is easier with underscores.
c2rcc_flags.Valid_PE && Lacul_Morii_Polygon
Thank you so much.
Can I bother you with one more question.
When trying to export the view to KMZ it requested the projection be made in Lat/Lon, since I forgot to make it in the first place, I tried to do it to the already c2rcc processed image but I get an error regarding Undefined symbol kz.
Can reprojection be made after c2rcc processing or it has to be done before processing?
Thank you!
It seems one of the expressions you are using is invalid. It uses the symbol kd_z90max_lulie. Is it a mask? Try to delete it.
Reprojection works after C2RCC. Not sure if C2RCC can handle the reprojected data.
Oh yes, I renamed some of the bands in order to better recognize them when using Statistics or Profile Plot. It worked by renaming them back into their original name. Now I can reproject all c2rcc processed images. Thank you!
1 Like