Dear Snap community,
I have a question relating to masking cloud cover when performing a biophysical processor to derive LAI at 10m by using S2A products.
Q1) Since I am using the S2A product in cloudy Ireland, will it be beneficial to further perform cloud correction before running the Biophysical processor, such as running the image through sen2cor software?
Q2) Is it best practice to perform a mask cover before or after performing the biophysical processor?
Q3) If we mask the cloud beforehand, then it is not clear which is the best algorithm to use, as it is not clear for the 10 m resolution derived LAI product, what is the maximum cloudiness (eg. opaque cloud or cirrus_cloud) the S2ToolBox can handle? Any recommendations on the methodology to mask clouds will be welcomed?
Many thanks for any help you may provide,
Joseph A.P. Pollacco
I can’t provide with a definite workflow, but I can give some hints. You can try and find out what works for you.
-
I think the LAI values or other results from the bio-physical processor will result in very specific values for cloudy pixels. Maybe you can use some threshold for the masking afterwards.
-
If this is not possible, you probably need to do it in parallel. A pre-generated mask can’t be used because the processor doesn’t support a user defined masking. So you need to compute the cloud mask and the bio-physical values in parallel, afterwards merge both and then you can use the mask bio-physical values.
-
For cloud screening I would suggest IdePix S2-MSI. You can find it among the available plugins. This generates a pixel_classif_flags band with several flags related to clouds.
-
I think there is no ‘maximum cloudiness’ the bio-physical processor can handle, because each value is calculated per pixel. So if there is only a single non-cloudy pixel in a scene it will be computed.
-
All can be realized by using a GPT Graph, once you know what you want to do. This can be applied to a batch of products.
Dear Marco,
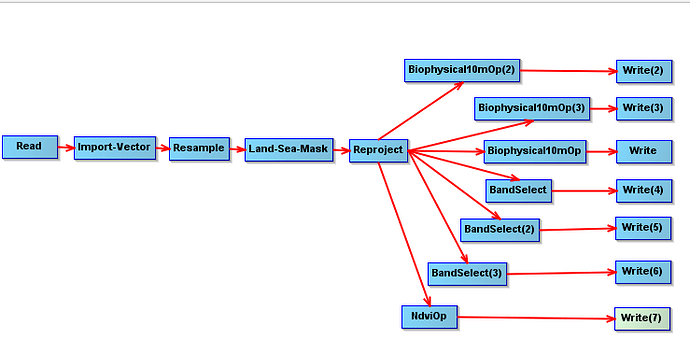
Greetings, and great thanks for putting me on the right path. Thanks to your recommendation, I successfully built a GPT Graph, for every satellite image, LAI, FAPAR, FVC (cover fraction), NDVI, OpaqueCloud, QualityCloudConfidence, and CirrusCloud are derived. I will have a time series, and cells with high cloud-cover values will be removed and interpolated from time series LAI (using Julia script) .
So the question is how to identify the problematic cells?
As you recommended, I tried using IdePix S2-MSI to derive pixel_classif_flags, but I was unable to use it because it requires Level-1C data.
Reading the manual, S2ToolBox Level 2 products: LAI, FAPAR, FCOVER, the bio-physical processor, as you pointed out, has a complex algorithm to handle cloud cover, so it may be best to contact S2ToolBox developers to see when they will fix their flag system.
I am wondering if, for example, questioning if QualityCloudConfidence threshold would be the simplest method of cloud screening until S2ToolBox gets updated?
With kindness,
Joseph A.P. Pollacco
Yes, the quality_cloud_confidence will already flag some of the cloudy pixels. As a placeholder till you have some better flag it will be sufficient. But this value is actually not good.
Also the values of the scene classification (scl) are not much better.
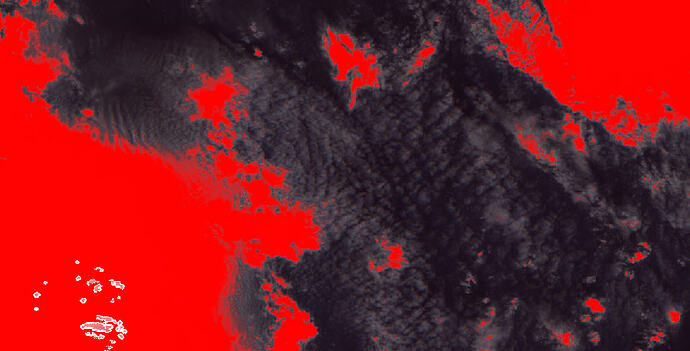
Attached is a snippet from an image in which I have overlaid the quality_cloud_confidence in red on an RGB. You still see a lot of remaining clouds.
Hi @Marco_EOM ,
A great thanks for your reply and for very kindly assisting with your testing.
I performed some tests with the Flags of the S2ToolBox of the Biophysical Processor which is Band 2 when saving for e.g. LAI.GeoTIFF.
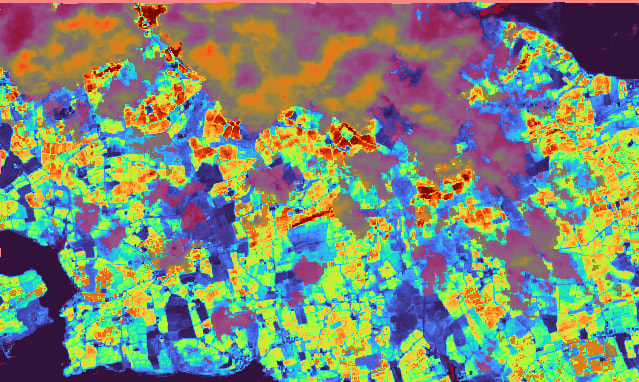
The Flags ranging from 1-17 and I found that the Flag=1 does a good job to mask the cloud of the LAI output.
Also I found this relationship by not using the Reprojection tool, which seemed to have caused the bugs.
Kind regards,
Joseph
The Flag=1 is semitransparent red of the LAI image is in Timoleague in Ireland:
1 Like