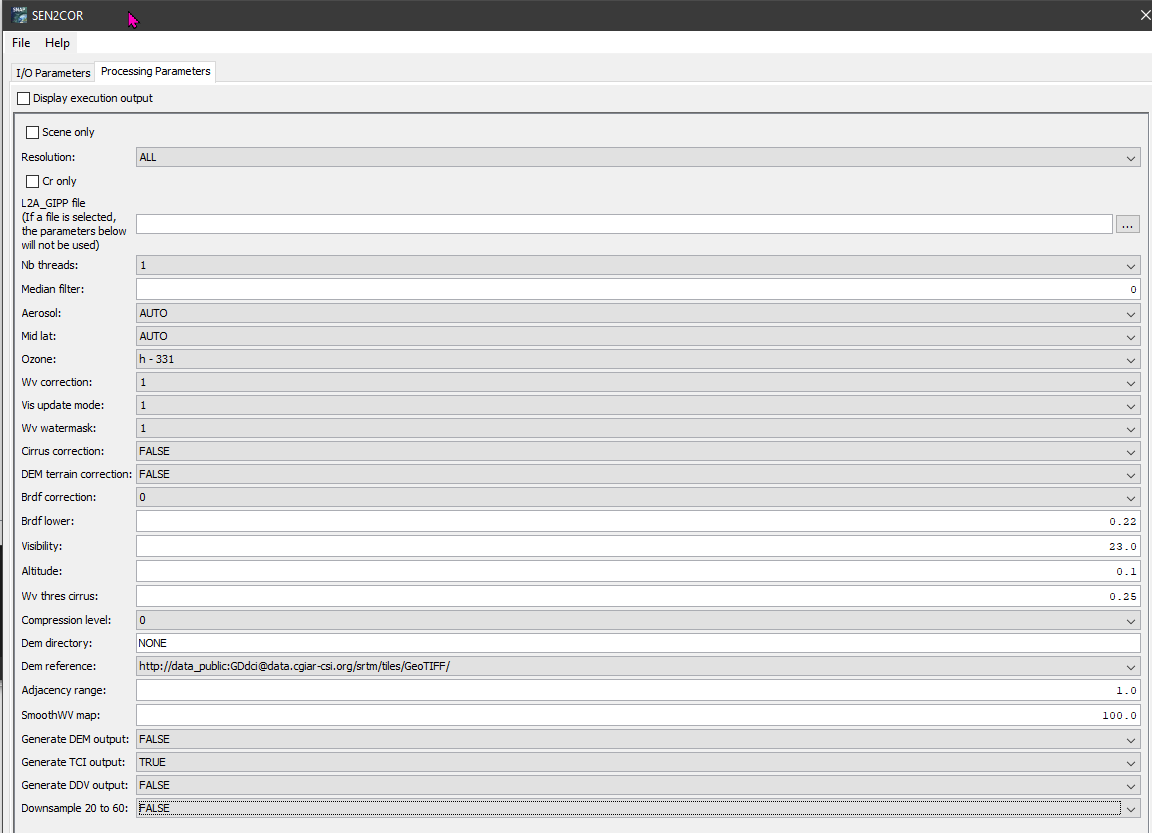
I was able to get the command line version to work but I am getting the same errors. I tried both 2.10 as well as the most recent version, 2.11:
Sen2Cor. Version: 02.10.01, created: 2021.12.13, supporting Level-1C product version 14.2 - 14.9 started …
Product version: 14.6
Operation mode: TOOLBOX
Processing baseline: 99.99
Progress[%]: 0.00 : Generating datastrip metadata
L2A datastrip successfully generated
No resolution specified, will process 20 and 10 m resolution
20 m resolution will be downsampled to 60 m
Progress[%]: 0.07 : PID-27904, L2A_ProcessTile: processing with resolution 20 m, elapsed time[s]: 1.720, total: 0:00:12.220000
Progress[%]: 0.07 : PID-27904, L2A_ProcessTile: start of pre processing, elapsed time[s]: 0.003, total: 0:00:12.223000
Progress[%]: 0.07 : PID-27904, L2A_Tables: start import, elapsed time[s]: 0.048, total: 0:00:12.271000
Progress[%]: 0.09 : PID-27904, L2A_Tables: band B01 imported, elapsed time[s]: 0.436, total: 0:00:12.707000
Progress[%]: 0.51 : PID-27904, L2A_Tables: band B02 imported, elapsed time[s]: 10.829, total: 0:00:23.536000
Progress[%]: 0.96 : PID-27904, L2A_Tables: band B03 imported, elapsed time[s]: 11.354, total: 0:00:34.890000
Progress[%]: 1.39 : PID-27904, L2A_Tables: band B04 imported, elapsed time[s]: 11.176, total: 0:00:46.066000
Progress[%]: 1.52 : PID-27904, L2A_Tables: band B05 imported, elapsed time[s]: 3.259, total: 0:00:49.325000
Progress[%]: 1.65 : PID-27904, L2A_Tables: band B06 imported, elapsed time[s]: 3.128, total: 0:00:52.453000
Progress[%]: 1.78 : PID-27904, L2A_Tables: band B07 imported, elapsed time[s]: 3.475, total: 0:00:55.928000
Progress[%]: 1.91 : PID-27904, L2A_Tables: band B8A imported, elapsed time[s]: 3.395, total: 0:00:59.323000
Progress[%]: 1.94 : PID-27904, L2A_Tables: band B09 imported, elapsed time[s]: 0.664, total: 0:00:59.987000
Progress[%]: 1.97 : PID-27904, L2A_Tables: band B10 imported, elapsed time[s]: 0.669, total: 0:01:00.656000
Progress[%]: 2.10 : PID-27904, L2A_Tables: band B11 imported, elapsed time[s]: 3.514, total: 0:01:04.170000
Progress[%]: 2.24 : PID-27904, L2A_Tables: band B12 imported, elapsed time[s]: 3.330, total: 0:01:07.500000
Progress[%]: 2.24 : PID-27904, L2A_Tables: AUX_CAMS* file not found., elapsed time[s]: 0.020, total: 0:01:07.520000
Progress[%]: 2.24 : PID-27904, L2A_Tables: No L1C AUX_DATA for aerosol optical thickness @ 550nm information, elapsed time[s]: 0.004, total: 0:01:07.524000
Progress[%]: 2.24 : PID-27904, L2A_Tables: CAMS ncfile: not present, elapsed time[s]: 0.005, total: 0:01:07.529000
Progress[%]: 2.24 : PID-27904, L2A_ProcessTile: start of Scene Classification, elapsed time[s]: 0.006, total: 0:01:07.535000
Progress[%]: 2.26 : PID-27904, L2A_Tables: band B02 must be resampled, elapsed time[s]: 0.696, total: 0:01:08.231000
Progress[%]: 2.41 : PID-27904, Pre process , elapsed time[s]: 3.721, total: 0:01:11.952000
Progress[%]: 2.41 : PID-27904, L2A_Tables: band B01 must be resampled, elapsed time[s]: 0.014, total: 0:01:11.966000
Progress[%]: 2.66 : PID-27904, L2A_Tables: band B03 must be resampled, elapsed time[s]: 6.361, total: 0:01:18.327000
Progress[%]: 2.81 : PID-27904, L2A_Tables: band B04 must be resampled, elapsed time[s]: 3.874, total: 0:01:22.201000
Progress[%]: 2.98 : PID-27904, L2A_Tables: band B09 must be resampled, elapsed time[s]: 4.191, total: 0:01:26.392000
Progress[%]: 3.17 : PID-27904, L2A_Tables: band B10 must be resampled, elapsed time[s]: 4.866, total: 0:01:31.258000
Progress[%]: 3.37 : PID-27904, L2A_SC init , elapsed time[s]: 5.156, total: 0:01:36.414000
Progress[%]: 3.40 : PID-27904, L2A_CSND_1_1 , elapsed time[s]: 0.703, total: 0:01:37.117000
Progress[%]: 3.42 : PID-27904, L2A_CSND_1_2 , elapsed time[s]: 0.692, total: 0:01:37.809000
Progress[%]: 3.43 : PID-27904, L2A_CSND_2_0 , elapsed time[s]: 0.199, total: 0:01:38.008000
Progress[%]: 3.46 : PID-27904, L2A_CSND_2_1 , elapsed time[s]: 0.664, total: 0:01:38.672000
Progress[%]: 3.47 : PID-27904, L2A_CSND_2_1_2, elapsed time[s]: 0.340, total: 0:01:39.012000
Progress[%]: 3.48 : PID-27904, L2A_CSND_2_2 , elapsed time[s]: 0.274, total: 0:01:39.286000
Progress[%]: 3.50 : PID-27904, L2A_CSND_2_3 , elapsed time[s]: 0.376, total: 0:01:39.662000
Progress[%]: 3.52 : PID-27904, L2A_CSND_2_4 , elapsed time[s]: 0.645, total: 0:01:40.307000
Progress[%]: 3.56 : PID-27904, L2A_CSND_2_5 , elapsed time[s]: 0.986, total: 0:01:41.293000
Progress[%]: 3.56 : PID-27904, L2A_SnowPostProcessingCCI , elapsed time[s]: 0.005, total: 0:01:41.298000
Progress[%]: 3.61 : PID-27904, L2A_CSND_3 , elapsed time[s]: 1.130, total: 0:01:42.428000
Progress[%]: 3.64 : PID-27904, L2A_CSND_5_1 , elapsed time[s]: 0.922, total: 0:01:43.350000
Progress[%]: 3.70 : PID-27904, L2A_CSND_5_2 , elapsed time[s]: 1.553, total: 0:01:44.903000
Progress[%]: 3.74 : PID-27904, L2A_CSND_6 , elapsed time[s]: 1.070, total: 0:01:45.973000
Progress[%]: 3.77 : PID-27904, L2A_CSND_6_2 , elapsed time[s]: 0.724, total: 0:01:46.697000
Progress[%]: 3.81 : PID-27904, L2A_CSND_7 , elapsed time[s]: 0.892, total: 0:01:47.589000
Progress[%]: 3.84 : PID-27904, DV recovery , elapsed time[s]: 0.920, total: 0:01:48.509000
Progress[%]: 3.88 : PID-27904, WP recovery , elapsed time[s]: 1.015, total: 0:01:49.524000
Progress[%]: 3.88 : PID-27904, WP recovery with CCI Water Bodies at 150m , elapsed time[s]: 0.005, total: 0:01:49.529000
Progress[%]: 3.93 : PID-27904, Snow recovery , elapsed time[s]: 1.095, total: 0:01:50.624000
Progress[%]: 4.07 : PID-27904, SnowSoilBorders, elapsed time[s]: 3.576, total: 0:01:54.200000
Progress[%]: 4.09 : PID-27904, Soil recovery , elapsed time[s]: 0.533, total: 0:01:54.733000
Progress[%]: 4.09 : PID-27904, Cirrus recovery with B10 , elapsed time[s]: 0.005, total: 0:01:54.738000
Progress[%]: 4.09 : PID-27904, Urban and Bare pixel recovery with CCI Land Cover Map at 300 m , elapsed time[s]: 0.004, total: 0:01:54.742000
Progress[%]: 4.41 : PID-27904, Bright pixels Hollstein Fmask, elapsed time[s]: 8.232, total: 0:02:02.974000
Cloud heights processed in 0:00:10.441000 seconds
Progress[%]: 6.03 : PID-27904, L2A_SHD , elapsed time[s]: 41.253, total: 0:02:44.227000
Progress[%]: 6.03 : PID-27904, WP recovery with CCI Water Bodies at 150m , elapsed time[s]: 0.005, total: 0:02:44.232000
Progress[%]: 6.07 : PID-27904, Topographic casted shadow Pixels post processing, elapsed time[s]: 1.146, total: 0:02:45.378000
Progress[%]: 6.18 : PID-27904, Post process , elapsed time[s]: 2.761, total: 0:02:48.139000
Progress[%]: 6.18 : PID-27904, L2A_ProcessTile: start of Atmospheric Correction, elapsed time[s]: 0.058, total: 0:02:48.197000
Progress[%]: 6.22 : PID-27904, L2A_AtmCorr: end of calculation terrain maps, elapsed time[s]: 0.852, total: 0:02:49.049000
Progress[%]: 6.22 : PID-27904, L2A_AtmCorr: start of AOT retrieval at 550nm, elapsed time[s]: 0.004, total: 0:02:49.053000
Progress[%]: 6.35 : PID-27904, L2A_AtmCorr: end of internal classification, elapsed time[s]: 3.386, total: 0:02:52.439000
Progress[%]: 6.36 : PID-27904, L2A_AtmCorr: end of interpolation LUTs, elapsed time[s]: 0.311, total: 0:02:52.750000
Progress[%]: 6.80 : PID-27904, L2A_AtmCorr: end retrieving reference pixels for dark areas, elapsed time[s]: 11.164, total: 0:03:03.914000
Progress[%]: 7.86 : PID-27904, L2A_AtmCorr: end of check for negative reflectance pixels, elapsed time[s]: 27.042, total: 0:03:30.956000
Progress[%]: 7.92 : PID-27904, L2A_AtmCorr: the rescaling of path radiance in blue band has been disabled by configuration, elapsed time[s]: 1.582, total: 0:03:32.538000
Progress[%]: 8.57 : PID-27904, L2A_AtmCorr: end of visibility index calculation, elapsed time[s]: 16.372, total: 0:03:48.910000
Progress[%]: 8.57 : PID-27904, L2A_AtmCorr: end of AOT retrieval at 550nm, elapsed time[s]: 0.005, total: 0:03:48.915000
Progress[%]: 8.57 : PID-27904, L2A_AtmCorr: start of water vapour retrieval, elapsed time[s]: 0.003, total: 0:03:48.918000
Progress[%]: 8.59 : PID-27904, L2A_AtmCorr: end of water vapour retrieval preparation, elapsed time[s]: 0.679, total: 0:03:49.597000
Progress[%]: 12.01 : PID-27904, L2A_AtmCorr: end of water vapour retrieval, elapsed time[s]: 87.263, total: 0:05:16.860000
Progress[%]: 12.01 : PID-27904, L2A_AtmCorr: preparation of surface reflectance retrieval, elapsed time[s]: 0.006, total: 0:05:16.866000
Progress[%]: 12.13 : PID-27904, L2A_AtmCorr: end of surface reflectance retrieval preparation, elapsed time[s]: 2.891, total: 0:05:19.757000
Progress[%]: 12.13 : PID-27904, L2A_AtmCorr: start of surface reflectance retrieval, elapsed time[s]: 0.004, total: 0:05:19.761000
Progress[%]: 16.86 : PID-27904, L2A_AtmCorr: end of surface reflectance retrieval, elapsed time[s]: 120.766, total: 0:07:20.527000
Progress[%]: 16.86 : PID-27904, L2A_AtmCorr: start of rho retrieval step 2, elapsed time[s]: 0.007, total: 0:07:20.534000
Progress[%]: 18.77 : PID-27904, L2A_AtmCorr: end of rho retrieval step 2, elapsed time[s]: 48.515, total: 0:08:09.049000
Progress[%]: 18.77 : PID-27904, L2A_ProcessTile: start of post processing, elapsed time[s]: 0.021, total: 0:08:09.070000
Progress[%]: 18.80 : PID-27904, L2A_Tables: start export for 20 m resolution, elapsed time[s]: 0.734, total: 0:08:09.804000
OpenJPEG function failure.
Traceback (most recent call last):
File “E:\Software\Sen2Cor-02.10.01-win64\Lib\site-packages\sen2cor\L2A_Tables.py”, line 3457, in exportBandList
self.glymurWrapper(filename, band)
File “E:\Software\Sen2Cor-02.10.01-win64\Lib\site-packages\sen2cor\L2A_Tables.py”, line 3617, in glymurWrapper
glymur.Jp2k(filename, band, **kwargs)
File “E:\Software\Sen2Cor-02.10.01-win64\lib\site-packages\glymur\jp2k.py”, line 177, in __init__
self._write(data, **kwargs)
File “E:\Software\Sen2Cor-02.10.01-win64\lib\site-packages\glymur\jp2k.py”, line 539, in _write
self._write_openjp2(img_array, verbose=verbose)
File “E:\Software\Sen2Cor-02.10.01-win64\lib\site-packages\glymur\jp2k.py”, line 783, in _write_openjp2
opj2.start_compress(codec, image, strm)
File “E:\Software\Sen2Cor-02.10.01-win64\lib\site-packages\glymur\lib\openjp2.py”, line 1299, in start_compress
OPENJP2.opj_start_compress(codec, image, stream)
File “E:\Software\Sen2Cor-02.10.01-win64\lib\site-packages\glymur\lib\openjp2.py”, line 594, in check_error
raise OpenJPEGLibraryError(“OpenJPEG function failure.”)
OpenJPEGLibraryError: OpenJPEG function failure.
Progress[%]: 18.89 : PID-27904, L2A_Tables: preparing downsampled export for 60 m resolution, elapsed time[s]: 2.282, total: 0:08:12.086000
Progress[%]: 18.89 : PID-27904, L2A_Tables: start export for 60 m resolution, elapsed time[s]: 0.005, total: 0:08:12.091000
no such child: Granule
Traceback (most recent call last):
File “E:\Software\Sen2Cor-02.10.01-win64\Lib\site-packages\sen2cor\L2A_Process.py”, line 692, in main
if tile.process() == True:
File “E:\Software\Sen2Cor-02.10.01-win64\Lib\site-packages\sen2cor\L2A_ProcessTileToolbox.py”, line 117, in process
if not self.process_resolution(20):
File “E:\Software\Sen2Cor-02.10.01-win64\Lib\site-packages\sen2cor\L2A_ProcessTileToolbox.py”, line 147, in process_resolution
return self._process()
File “E:\Software\Sen2Cor-02.10.01-win64\Lib\site-packages\sen2cor\L2A_ProcessTileToolbox.py”, line 197, in _process
if self.postprocess() == False:
File “E:\Software\Sen2Cor-02.10.01-win64\Lib\site-packages\sen2cor\L2A_ProcessTileToolbox.py”, line 273, in postprocess
res = self.tables.downsampleBandList_20to60_andExport()
File “E:\Software\Sen2Cor-02.10.01-win64\Lib\site-packages\sen2cor\L2A_Tables.py”, line 3096, in downsampleBandList_20to60_andExport
Granule = pi.Product_Organisation.Granule_List.Granule
File “src/lxml/lxml.objectify.pyx”, line 229, in lxml.objectify.ObjectifiedElement.__getattr__ (src\lxml\lxml.objectify.c:3835)
File “src/lxml/lxml.objectify.pyx”, line 450, in lxml.objectify._lookupChildOrRaise (src\lxml\lxml.objectify.c:6539)
AttributeError: no such child: Granule
Progress[%]: 100.00 : Application terminated with at least one error.
_______________________________________________________________________
Sen2Cor. Version: 02.11.00, created: 2022.10.20, supporting Level-1C product version 14.2 - 14.9 started …
Product version: 14.6
Operation mode: TOOLBOX
Processing baseline: 99.99
Progress[%]: 0.00 : Generating datastrip metadata
L2A datastrip successfully generated
No resolution specified, will process 20 and 10 m resolution
20 m resolution will be downsampled to 60 m
Progress[%]: 0.13 : PID-13576, L2A_ProcessTile: processing with resolution 20 m, elapsed time[s]: 3.241, total: 0:00:14.892000
Progress[%]: 0.13 : PID-13576, L2A_ProcessTile: start of pre processing, elapsed time[s]: 0.004, total: 0:00:14.896000
Progress[%]: 0.15 : PID-13576, L2A_Tables: start import, elapsed time[s]: 0.488, total: 0:00:15.384000
Progress[%]: 0.18 : PID-13576, L2A_Tables: band B01 imported, elapsed time[s]: 0.800, total: 0:00:16.184000
Progress[%]: 0.62 : PID-13576, L2A_Tables: band B02 imported, elapsed time[s]: 11.211, total: 0:00:27.395000
Progress[%]: 1.11 : PID-13576, L2A_Tables: band B03 imported, elapsed time[s]: 12.685, total: 0:00:40.080000
Progress[%]: 1.63 : PID-13576, L2A_Tables: band B04 imported, elapsed time[s]: 13.032, total: 0:00:53.112000
Progress[%]: 1.78 : PID-13576, L2A_Tables: band B05 imported, elapsed time[s]: 3.906, total: 0:00:57.018000
Progress[%]: 1.93 : PID-13576, L2A_Tables: band B06 imported, elapsed time[s]: 3.759, total: 0:01:00.777000
Progress[%]: 2.09 : PID-13576, L2A_Tables: band B07 imported, elapsed time[s]: 4.158, total: 0:01:04.935000
Progress[%]: 2.26 : PID-13576, L2A_Tables: band B8A imported, elapsed time[s]: 4.262, total: 0:01:09.197000
Progress[%]: 2.28 : PID-13576, L2A_Tables: band B09 imported, elapsed time[s]: 0.680, total: 0:01:09.877000
Progress[%]: 2.31 : PID-13576, L2A_Tables: band B10 imported, elapsed time[s]: 0.772, total: 0:01:10.649000
Progress[%]: 2.49 : PID-13576, L2A_Tables: band B11 imported, elapsed time[s]: 4.453, total: 0:01:15.102000
Progress[%]: 2.65 : PID-13576, L2A_Tables: band B12 imported, elapsed time[s]: 4.191, total: 0:01:19.293000
Progress[%]: 2.66 : PID-13576, L2A_Tables: AUX_CAMS* file not found., elapsed time[s]: 0.214, total: 0:01:19.507000
Progress[%]: 2.66 : PID-13576, L2A_Tables: No L1C AUX_DATA for aerosol optical thickness @ 550nm information, elapsed time[s]: 0.004, total: 0:01:19.511000
Progress[%]: 2.66 : PID-13576, L2A_Tables: CAMS ncfile: not present, elapsed time[s]: 0.006, total: 0:01:19.517000
Progress[%]: 2.66 : PID-13576, L2A_ProcessTile: start of Scene Classification, elapsed time[s]: 0.006, total: 0:01:19.523000
Progress[%]: 2.87 : PID-13576, L2A_Tables: band B02 must be resampled, elapsed time[s]: 5.280, total: 0:01:24.803000
Progress[%]: 3.02 : PID-13576, Pre process , elapsed time[s]: 3.736, total: 0:01:28.539000
Progress[%]: 3.02 : PID-13576, L2A_Tables: band B01 must be resampled, elapsed time[s]: 0.015, total: 0:01:28.554000
Progress[%]: 3.29 : PID-13576, L2A_Tables: band B03 must be resampled, elapsed time[s]: 6.918, total: 0:01:35.472000
Progress[%]: 3.43 : PID-13576, L2A_Tables: band B04 must be resampled, elapsed time[s]: 3.705, total: 0:01:39.177000
Progress[%]: 3.60 : PID-13576, L2A_Tables: band B09 must be resampled, elapsed time[s]: 4.257, total: 0:01:43.434000
Progress[%]: 3.79 : PID-13576, L2A_Tables: band B10 must be resampled, elapsed time[s]: 4.969, total: 0:01:48.403000
Progress[%]: 4.00 : PID-13576, L2A_SC init , elapsed time[s]: 5.214, total: 0:01:53.617000
Progress[%]: 4.03 : PID-13576, L2A_CSND_1_1 , elapsed time[s]: 0.760, total: 0:01:54.377000
Progress[%]: 4.06 : PID-13576, L2A_CSND_1_2 , elapsed time[s]: 0.780, total: 0:01:55.157000
Progress[%]: 4.13 : PID-13576, L2A_CSND_2_0 , elapsed time[s]: 1.711, total: 0:01:56.868000
Progress[%]: 4.16 : PID-13576, L2A_CSND_2_1 , elapsed time[s]: 0.746, total: 0:01:57.614000
Progress[%]: 4.17 : PID-13576, L2A_CSND_2_1_2, elapsed time[s]: 0.380, total: 0:01:57.994000
Progress[%]: 4.18 : PID-13576, L2A_CSND_2_2 , elapsed time[s]: 0.317, total: 0:01:58.311000
Progress[%]: 4.20 : PID-13576, L2A_CSND_2_3 , elapsed time[s]: 0.411, total: 0:01:58.722000
Progress[%]: 4.23 : PID-13576, L2A_CSND_2_4 , elapsed time[s]: 0.691, total: 0:01:59.413000
Progress[%]: 4.27 : PID-13576, L2A_CSND_2_5 , elapsed time[s]: 1.050, total: 0:02:00.463000
Progress[%]: 4.27 : PID-13576, L2A_SnowPostProcessingCCI , elapsed time[s]: 0.005, total: 0:02:00.468000
Progress[%]: 4.32 : PID-13576, L2A_CSND_3 , elapsed time[s]: 1.253, total: 0:02:01.721000
Progress[%]: 4.36 : PID-13576, L2A_CSND_5_1 , elapsed time[s]: 1.069, total: 0:02:02.790000
Progress[%]: 4.42 : PID-13576, L2A_CSND_5_2 , elapsed time[s]: 1.657, total: 0:02:04.447000
Progress[%]: 4.47 : PID-13576, L2A_CSND_6 , elapsed time[s]: 1.194, total: 0:02:05.641000
Progress[%]: 4.50 : PID-13576, L2A_CSND_6_2 , elapsed time[s]: 0.775, total: 0:02:06.416000
Progress[%]: 4.54 : PID-13576, L2A_CSND_7 , elapsed time[s]: 0.935, total: 0:02:07.351000
Progress[%]: 4.58 : PID-13576, DV recovery , elapsed time[s]: 0.989, total: 0:02:08.340000
Progress[%]: 4.62 : PID-13576, WP recovery , elapsed time[s]: 1.108, total: 0:02:09.448000
Progress[%]: 4.62 : PID-13576, WP recovery with CCI Water Bodies at 150m , elapsed time[s]: 0.005, total: 0:02:09.453000
Progress[%]: 4.66 : PID-13576, Snow recovery , elapsed time[s]: 1.120, total: 0:02:10.573000
Progress[%]: 4.80 : PID-13576, SnowSoilBorders, elapsed time[s]: 3.583, total: 0:02:14.156000
Progress[%]: 4.83 : PID-13576, Soil recovery , elapsed time[s]: 0.562, total: 0:02:14.718000
Progress[%]: 4.83 : PID-13576, Cirrus recovery with B10 , elapsed time[s]: 0.006, total: 0:02:14.724000
Progress[%]: 4.83 : PID-13576, Urban and Bare pixel recovery with CCI Land Cover Map at 300 m , elapsed time[s]: 0.004, total: 0:02:14.728000
Progress[%]: 5.16 : PID-13576, Bright pixels Hollstein Fmask, elapsed time[s]: 8.438, total: 0:02:23.166000
Cloud heights processed in 0:00:10.989000 seconds
Progress[%]: 6.93 : PID-13576, L2A_SHD , elapsed time[s]: 45.179, total: 0:03:08.345000
Progress[%]: 6.93 : PID-13576, WP recovery with CCI Water Bodies at 150m , elapsed time[s]: 0.006, total: 0:03:08.351000
Progress[%]: 6.97 : PID-13576, Topographic casted shadow Pixels post processing, elapsed time[s]: 1.141, total: 0:03:09.492000
Progress[%]: 7.10 : PID-13576, Post process , elapsed time[s]: 3.133, total: 0:03:12.625000
Progress[%]: 7.10 : PID-13576, L2A_ProcessTile: start of Atmospheric Correction, elapsed time[s]: 0.060, total: 0:03:12.685000
Progress[%]: 7.14 : PID-13576, L2A_AtmCorr: end of calculation terrain maps, elapsed time[s]: 0.962, total: 0:03:13.647000
Progress[%]: 7.14 : PID-13576, L2A_AtmCorr: start of AOT retrieval at 550nm, elapsed time[s]: 0.017, total: 0:03:13.664000
Progress[%]: 7.27 : PID-13576, L2A_AtmCorr: end of internal classification, elapsed time[s]: 3.358, total: 0:03:17.022000
Progress[%]: 7.28 : PID-13576, L2A_AtmCorr: end of interpolation LUTs, elapsed time[s]: 0.310, total: 0:03:17.332000
Progress[%]: 7.72 : PID-13576, L2A_AtmCorr: end retrieving reference pixels for dark areas, elapsed time[s]: 11.204, total: 0:03:28.536000
Progress[%]: 8.79 : PID-13576, L2A_AtmCorr: end of check for negative reflectance pixels, elapsed time[s]: 27.160, total: 0:03:55.696000
Progress[%]: 8.85 : PID-13576, L2A_AtmCorr: the rescaling of path radiance in blue band has been disabled by configuration, elapsed time[s]: 1.634, total: 0:03:57.330000
Progress[%]: 9.51 : PID-13576, L2A_AtmCorr: end of visibility index calculation, elapsed time[s]: 16.740, total: 0:04:14.070000
Progress[%]: 9.51 : PID-13576, L2A_AtmCorr: end of AOT retrieval at 550nm, elapsed time[s]: 0.005, total: 0:04:14.075000
Progress[%]: 9.51 : PID-13576, L2A_AtmCorr: start of water vapour retrieval, elapsed time[s]: 0.004, total: 0:04:14.079000
Progress[%]: 9.53 : PID-13576, L2A_AtmCorr: end of water vapour retrieval preparation, elapsed time[s]: 0.658, total: 0:04:14.737000
Progress[%]: 13.02 : PID-13576, L2A_AtmCorr: end of water vapour retrieval, elapsed time[s]: 88.946, total: 0:05:43.683000
Progress[%]: 13.02 : PID-13576, L2A_AtmCorr: preparation of surface reflectance retrieval, elapsed time[s]: 0.005, total: 0:05:43.688000
Progress[%]: 13.13 : PID-13576, L2A_AtmCorr: end of surface reflectance retrieval preparation, elapsed time[s]: 2.880, total: 0:05:46.568000
Progress[%]: 13.13 : PID-13576, L2A_AtmCorr: start of surface reflectance retrieval, elapsed time[s]: 0.004, total: 0:05:46.572000
Progress[%]: 17.86 : PID-13576, L2A_AtmCorr: end of surface reflectance retrieval, elapsed time[s]: 120.400, total: 0:07:46.972000
Progress[%]: 17.86 : PID-13576, L2A_AtmCorr: start of rho retrieval step 2, elapsed time[s]: 0.012, total: 0:07:46.984000
Progress[%]: 19.76 : PID-13576, L2A_AtmCorr: end of rho retrieval step 2, elapsed time[s]: 48.670, total: 0:08:35.654000
Progress[%]: 19.77 : PID-13576, L2A_ProcessTile: start of post processing, elapsed time[s]: 0.022, total: 0:08:35.676000
Progress[%]: 19.79 : PID-13576, L2A_Tables: start export for 20 m resolution, elapsed time[s]: 0.738, total: 0:08:36.414000
OpenJPEG function failure.
Traceback (most recent call last):
File “E:\Software\Sen2Cor-02.11.00-win64\Lib\site-packages\sen2cor\L2A_Tables.py”, line 3543, in exportBandList
self.glymurWrapper(filename, band)
File “E:\Software\Sen2Cor-02.11.00-win64\Lib\site-packages\sen2cor\L2A_Tables.py”, line 3703, in glymurWrapper
glymur.Jp2k(filename, band, **kwargs)
File “E:\Software\Sen2Cor-02.11.00-win64\lib\site-packages\glymur\jp2k.py”, line 177, in __init__
self._write(data, **kwargs)
File “E:\Software\Sen2Cor-02.11.00-win64\lib\site-packages\glymur\jp2k.py”, line 539, in _write
self._write_openjp2(img_array, verbose=verbose)
File “E:\Software\Sen2Cor-02.11.00-win64\lib\site-packages\glymur\jp2k.py”, line 783, in _write_openjp2
opj2.start_compress(codec, image, strm)
File “E:\Software\Sen2Cor-02.11.00-win64\lib\site-packages\glymur\lib\openjp2.py”, line 1299, in start_compress
OPENJP2.opj_start_compress(codec, image, stream)
File “E:\Software\Sen2Cor-02.11.00-win64\lib\site-packages\glymur\lib\openjp2.py”, line 594, in check_error
raise OpenJPEGLibraryError(“OpenJPEG function failure.”)
OpenJPEGLibraryError: OpenJPEG function failure.
Progress[%]: 19.88 : PID-13576, L2A_Tables: preparing downsampled export for 60 m resolution, elapsed time[s]: 2.194, total: 0:08:38.608000
Progress[%]: 19.88 : PID-13576, L2A_Tables: start export for 60 m resolution, elapsed time[s]: 0.005, total: 0:08:38.613000
no such child: Granule
Traceback (most recent call last):
File “E:\Software\Sen2Cor-02.11.00-win64\Lib\site-packages\sen2cor\L2A_Process.py”, line 692, in main
if tile.process() == True:
File “E:\Software\Sen2Cor-02.11.00-win64\Lib\site-packages\sen2cor\L2A_ProcessTileToolbox.py”, line 117, in process
if not self.process_resolution(20):
File “E:\Software\Sen2Cor-02.11.00-win64\Lib\site-packages\sen2cor\L2A_ProcessTileToolbox.py”, line 147, in process_resolution
return self._process()
File “E:\Software\Sen2Cor-02.11.00-win64\Lib\site-packages\sen2cor\L2A_ProcessTileToolbox.py”, line 197, in _process
if self.postprocess() == False:
File “E:\Software\Sen2Cor-02.11.00-win64\Lib\site-packages\sen2cor\L2A_ProcessTileToolbox.py”, line 273, in postprocess
res = self.tables.downsampleBandList_20to60_andExport()
File “E:\Software\Sen2Cor-02.11.00-win64\Lib\site-packages\sen2cor\L2A_Tables.py”, line 3156, in downsampleBandList_20to60_andExport
Granule = pi.Product_Organisation.Granule_List.Granule
File “src/lxml/lxml.objectify.pyx”, line 229, in lxml.objectify.ObjectifiedElement.__getattr__ (src\lxml\lxml.objectify.c:3835)
File “src/lxml/lxml.objectify.pyx”, line 450, in lxml.objectify._lookupChildOrRaise (src\lxml\lxml.objectify.c:6539)
AttributeError: no such child: Granule
Progress[%]: 100.00 : Application terminated with at least one error.
![]()