Hi,everyone,
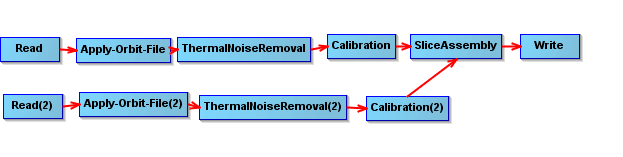
I wanted to preprocess the Sentinel-1 GRD data and chose to use the GraphBuilder and Batchprocessing tool in the GUI of SNAP.I created the processing chain as shown follow .However, the Slice Assembly tool didn’t work well in Batchprocessing as the two images I input didn’t get assembled.What should I do ?
Regards,
Wendy
Hi @WenjiaYan,
you may have to check if Thermal noise removal work on the whole SLC product at once (including all the subswaths), if not then you have to use TOPS split to separate them and then apply the Thermal noise removal to individual sub swaths.
Best
I would to Slice Assembly first since to me that is the “lowest-level” operation of reconstructing the original data-take.
Hello,@waqas782@mengdahl
I’m so sorry that I wrote the wrong in product type.It’s GRDH product.This processing chain can work well in the GraphBuilder,but in
the BatchProcessing I’m confused about how to deal with the input images since the Slice assembly tool need a pair of images . The Find-pair-image tool may be useful ,I don’t know how to employ it .
Thanks for the help!
Wendy
The batch processing only supports processing a list of products for a graph with a single input product and a single output product.
It should be possible to store the graph to disk and then run it on the command line with gpt. There you don’t have the one to one limitation as in the batch processing GUI.
Hello marpet,
I have the same question with @WenjiaYan . Can you tell me how to deal with the problem with GPT on the command line ?
Thank you !
there is the tool called gpt. You can find it in the bin folder of the SNAP installation directory.
On Windows you have the shortcut in the start menu ‘SNAP Command-Line’. There you can invoke gpt.
Call gpt -h to get help.
There is a tutorial on command line processing on the tutorials page http://step.esa.int/main/doc/tutorials/
Hello,developers
Thanks for all replies. I read the tutorial on command line processing and I tried to create the command line with the following codes in figure1.There is an error:[NodeId:Read] Specified ‘file’ does not exist.
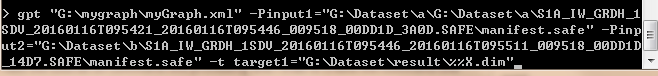 figure1
figure1
the graph file is as follow:
myGraph.xml (4.9 KB)
I will appreciate any helps!
Wendy
Probably because of the path you use:
G:\Datatset\a\G:\Dataset\a…
Thank you very much!I tried again and another error is that :I/O error reading image metadata.
Best regards,
Wendy
Can you run it again with the option -e.
With this option you should get more information about the error.
Please post the full error message.
Hi,marpet
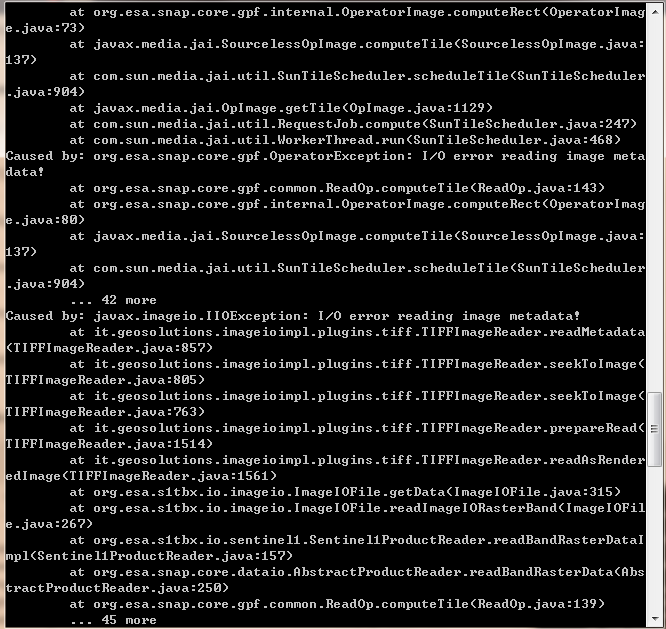
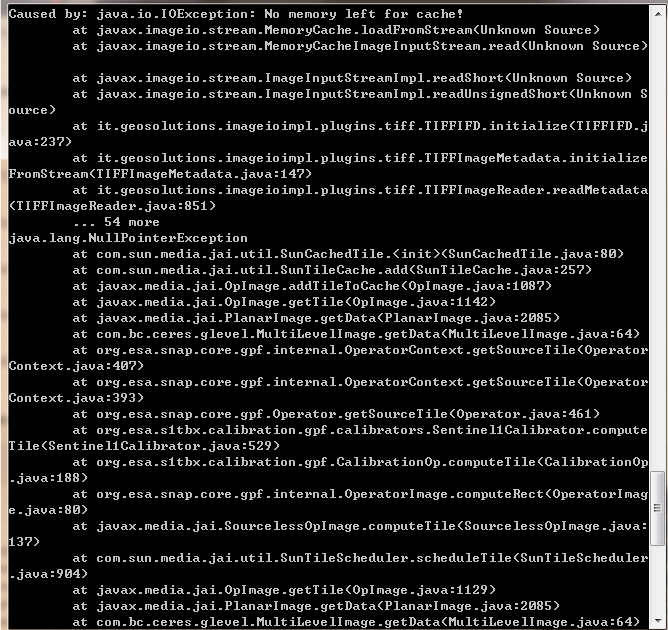
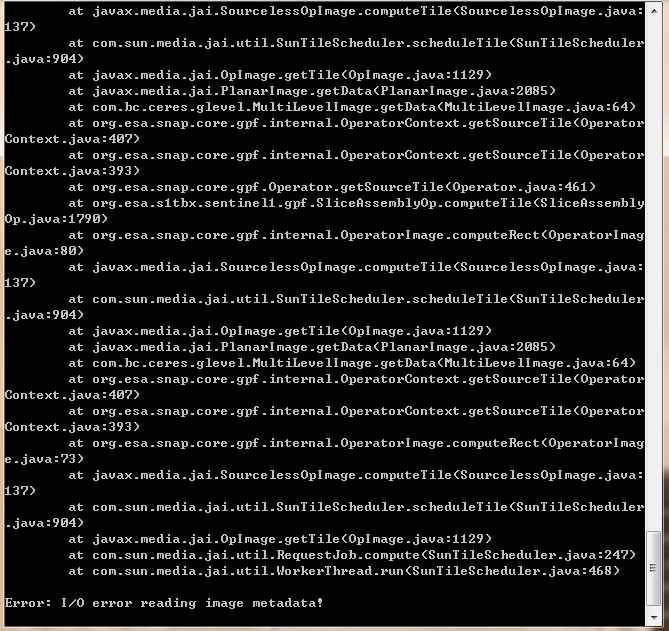
I ran it with the option -e and the error message is as follows:
I’m so sorry to bother you!
Best,
Wendy
How much memory do you have?
Because of “No memory left for cache!”, I think you don’t have enough memory.
The CPU is 8G.I’m confused about whether there are any mistakes in my command line statement.
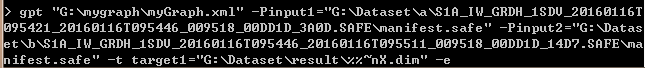
Thank you very much!
Wendy
No, I don’t think that it is caused by your command. It looks good. Except the target file name.
It seems the variable %%~nX is not properly resolved. But I don’t think that this causes error.
@lveci The error occurs within your Sentinel1ProductReader. Have you seen it before?
Maybe this post from Luis helps:
Hello,marpet
I really appreciate your helping me !I checked the option . There is no error this time but no output from the command.What’s wrong?
Regards,
Wendy
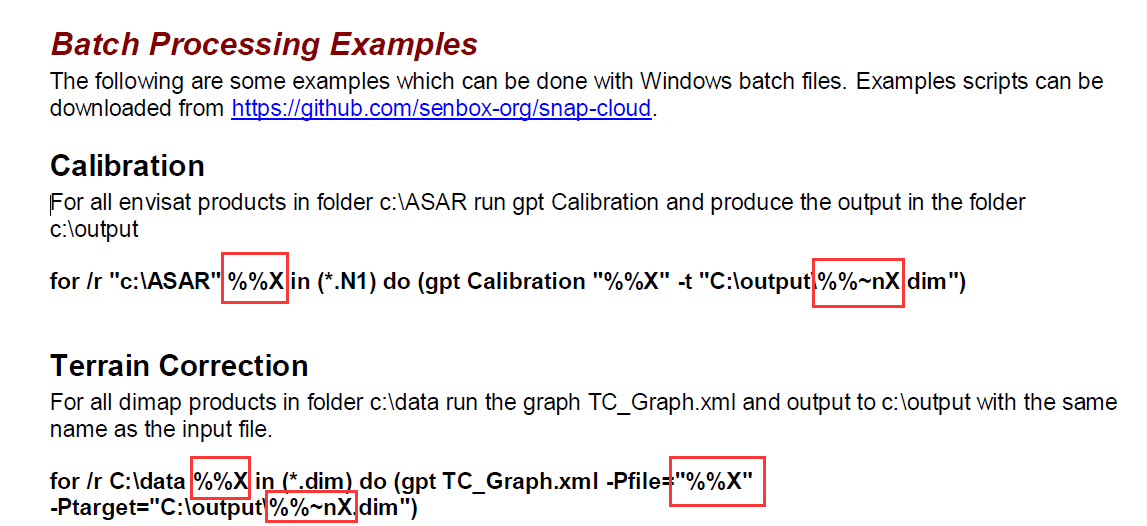
Hi, @lveci
I’m so sorry to bother you since I really need your help. I’m confused about the expressions in the red rectangle of the figure1 and what’re their meanings ,just like regular expression? In addition, is it possible to use the for-command to go through two folders with images which are processed by Slice Assembly ?If ok, how to do it ?
Thank you for any help!
Wendy