Hi all,
I am having issues with loading Sentinel-2 data into SNAP. I’m kind of new to this and haven’t used SNAP before but I guess you can just kind of load a downloaded zip file into the software. However, I tried this on SNAP 13 (uninstalled and re-installed) and SNAP 12, but none of them would load the imagery. Steps that I follow:
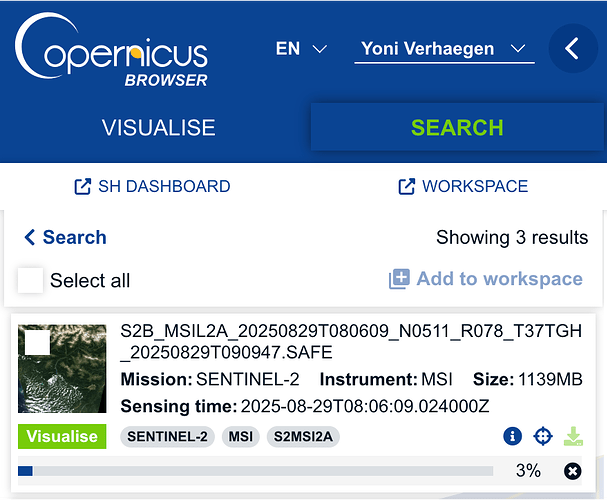
- download the data from Copernicus Browser:
which gives me:
and in the end the zip file is just in my folder like:
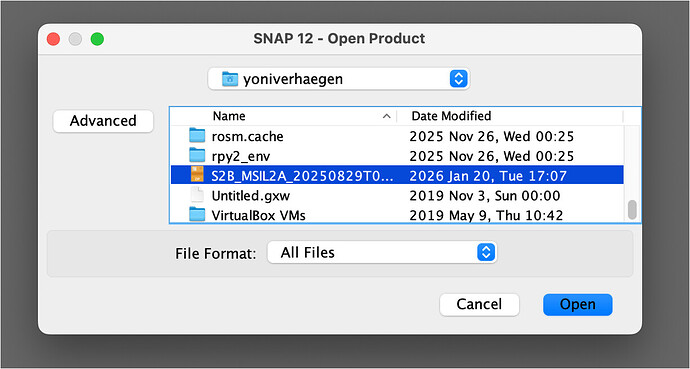
- I try to load it into SNAP (here version 12) :
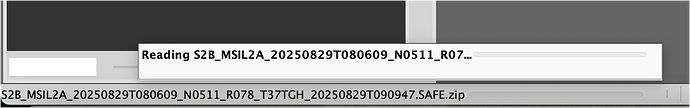
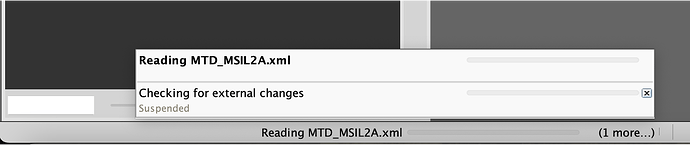
which just gives me this, no progression at all:
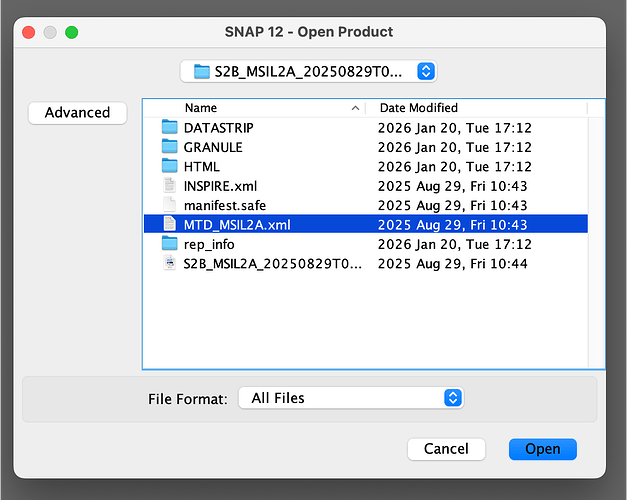
- then I tried unzipping the file and load the .xml:
same thing, nothing happens:
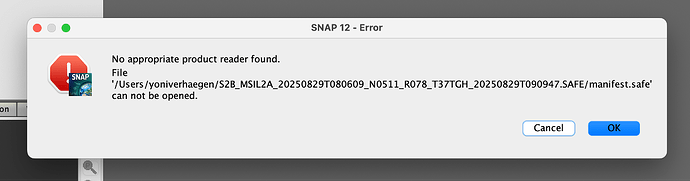
and with the manifest.safe file:
Can someone give me any guidance on this, it shouldn’t be that hard just loading in a file, right? Anyway, I do not proceed in doing so.
Thanks!
Hi @yoni.verhaegen
Could you please check whether the following directory has been created and attach a screenshot of its contents?
/Users/<user>/.snap/auxdata/gdal
Also, please check whether there is a gdal.lock file at the same level as the gdal folder. If you find it, please delete it and restart SNAP.
Thank you,
Diana
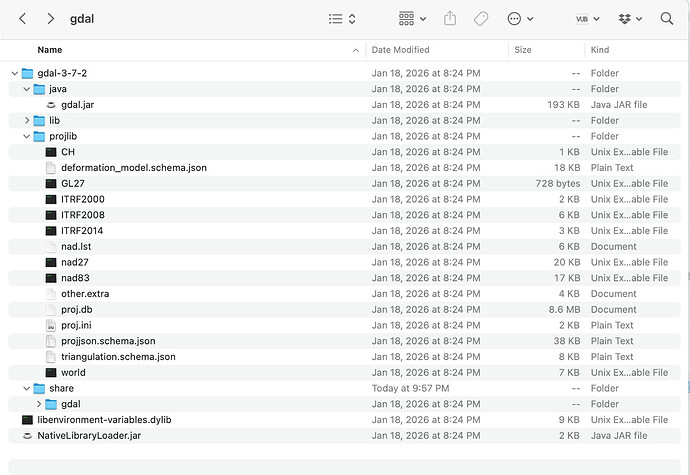
Hi! Thanks for the response, the folder looks like this:
see comment above. I also deleted the lock file but didn’t solve the problem. In fact, every time I try to load in the .zip file the lock file appears again.
Could you try deleting the entire gdal folder as well?
Also, could you provide some information about your OS?
Thank you!
Hi, see above for the OS info  deleting the gdal folder does not work…
deleting the gdal folder does not work…
Did you use the macOS ARM or the macOS Intel installer?
We tested on macOS Monterey version 12.7.4 and could not reproduce the issue.
Could you please attach the messages.log file generated after reproducing the issue?
Thank you,
Diana
Hello @diana_harosa
A lot of users have the same problems with Sentinel-2 products on macOS, I was looking for help too, and different forum discussions about solutions were all heavily programming related and not 100% guaranteed to work. This error seems to occur since around 2022.
Sentinel-1 data works, but Sen2 always ends up with these Errors. SNAP 13.0 version, macOS Tahoe 26.4.1
Thank you for your time in advance.
Hello @iceicefelix ,
Could you please attach the log file generated after reproducing the error?
Thank you!
Hi, sure. @diana_harosa
messages.log (191.9 KB)
I also have another error occurring, when trying to open a band in image view:
“Failed to open image view.
expression: Undefined symbol ‘WQSF_Isb.OC4ME_FAIL’.”
“
This time its with Sen3 OLCI, but have it on Sen1 as well occurring.
The product got many WQSF.lsb flags, but no with the _OC4ME_FAIL ending.
It might be a problem on my line of understanding, but I would really appreciate you looking into that. I’d love to work more with SNAP data instead of lesser available GIS capable data.
Edit: Error occurs due to the New Sentinel-3 OLCI level 2 Water processing update.The OC4ME band isn’t available in data after 26.02.2026, which flags the error. When I use data previous to that date opening images works.
Hi Felix,
There is a new reprocessing of Sentinel-3 OLCI (BC004) and there has been a change in the algorithm for chlorophyll estimation. The OC4ME does not exist anymore in the new reprocessing. Probably you are mixing old and new version of OLCI: EUMETSAT - User Portal
The old reader is not able to find the OC4ME flag (Renamed: OC4ME_FAIL → CHLOR_A_FAIL) and that is why it fails. Have you updated SNAP v.13 lately?