Hi
I uses sen2core in snap to process sentinel 2 level 1c to level 2a with sen2core this data i got from scihub.esa . but result of sen2core can’t open maybe someone here could help me.
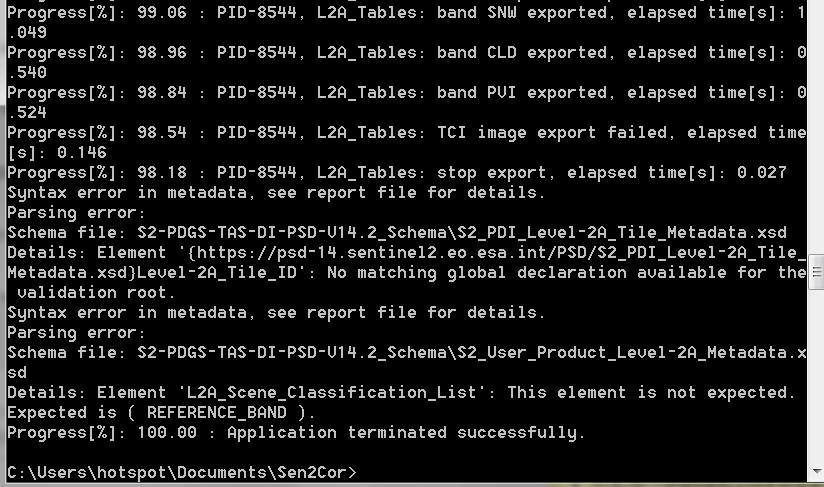
btw i used cmd prompt to process sen2core here is the screenshoot
and here the result
i dont know if my process succes or not but there is a new folder which one that folder contain metadata named MTD_MSIL2A.XML but it cant opened
please help if anyone know the answer
What versions of Sen2Cor and SNAP are you using?
i using snap 6.0 and sen2cor 2.4.0.when i trace the cmd i think there is something with the process so it’s cause made the outoput not finished yet. could you suggest me what should i do?did 1 should download data 1c again or download the sen2core again?
or i have problem with extract the rar?
thanks
What error do you have when you open the product in SNAP?
In the folder S2A_MSIL2A_20180219T025751_N0206_R032_T48MYT_20180219T073142.SAFE\GRANULE\L2A_T48MYT_A013900_20180219T031153\IMG_DATA\ have been generated the expected jp2 images?
Do you have problems only with this product?
@obarrilero thanks for your attention before.there is no jp2 in folder that you mean. I have already read another forum about using 7.zip and put on the folder which want to process in c/user document and it works.so now i can open process from sen 2 core.
thanks