hi everyone
i’m first time using SNAP and Sentinel 2 data . i have to create subset of the image for this purpose i am re sampling the image but the function is creating error . i read some similar topics related to this problem but problem is still present. i need help to solve this problem.
thanks in advance.
How much RAM does your system have? Maybe it is not enough.
I have 12 GB RAM in my system.
@marpet thanks for reply
please also tell me the basic requirements of the system to operate S2 data completely.
This always depends on what you are doing with the data. How many and which operations you are using in your processing chains.
I’m having 16GB and I can do almost everything with Sentinel-2 data.
i wanted to subset S2 image , but it required resampling first i use almost all types of operations for resampling but every time is error occur. now what you think , what should do to solve this problem.
I just tried your use case. I resampled to 10m and tailored the result to a region of 4000x4000 pixels.
I’ve limmited SNAP not to use more than 8GB of RAM and it worked smoothly.
Actually, SNAP never used more than 5GB during the process.
I guess that your SNAP is configured by default also to use 8GB.
So, you could try to increase this value.
In the ‘etc’ folder of the SNAP installation directory you’ll find a file named snap.conf. Open it in a text editor.
There is the line which starts with ‘default_options=’
In this line you’ll find an option like -J-Xmx8G. Increase the value. You could use something like -J-Xmx10G.
If you experience the error on the command line with gpt or pconvert you need to change different files. You need to change the corresponding vmoptions files, either gpt.vmoptions or pconvert.vmoptions. Change the value after -Xmx in the last line.
@marpet thank you for reply. i follow your instruction and go to C:\Users\arehm.snap\etc but i does’t found the file with name snap.conf only three readable files are available as you see in the the pic that i attached with this.
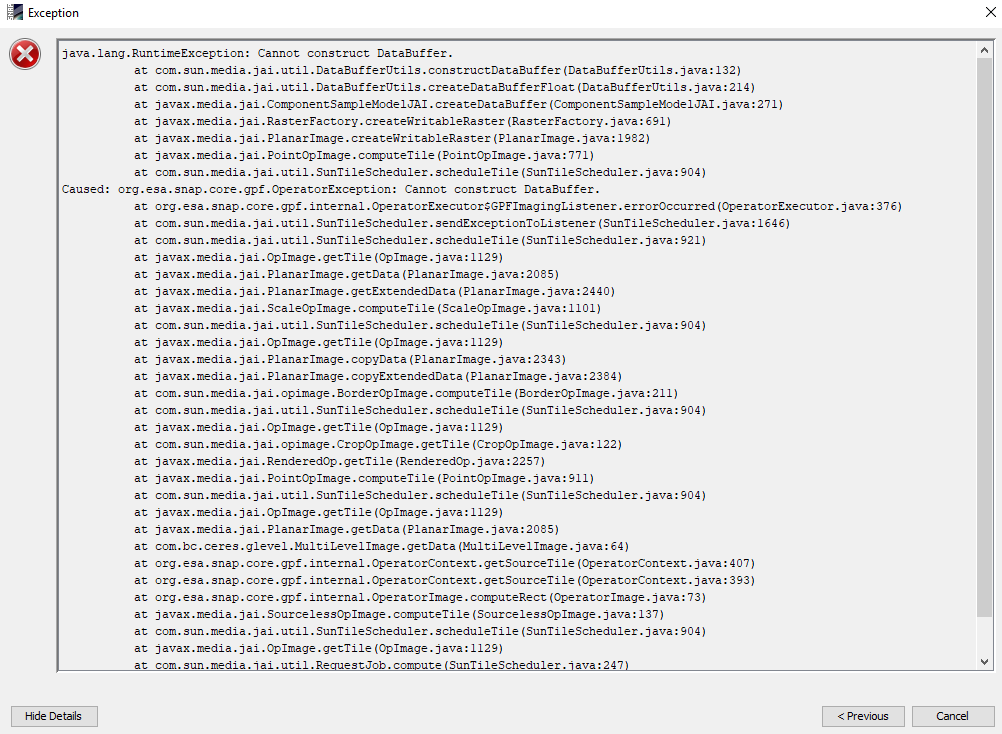
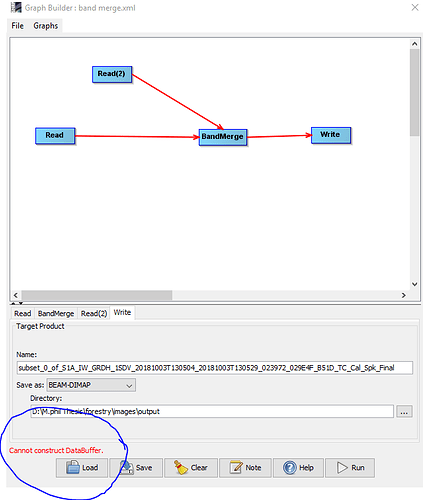
this error is not only in resampling S2 data. i’m also working on S1 data , when i try to do band marge databuffer also create
.is any possibility to operate SNAP smoothly just like other software like ERDAS
This is the snap user directory you found. 
I was referring to the installation directory.
It should be C:\Program Files\snap
i found that file and change J-Xmx7G To J-Xmx10G but the file does not saved . i restart my laptop and do this again but file does note save .
I also try to change this in SNAP tools>options>performance but it is not editable.
This is caused by the location of these files under C:\Program Files\ which is protected from editing.
Open the files as an administrator as described below to edit and save them.
thanks @marpet and @ABraun for guideline. problem has solved i just increase RAM from 12 to 16 GB and reinstall SNAP.
now it’s all function working properly.
Hello, I’m facing the same problem when processing my recently downloaded S2 files, I’ve tried to proceed it using a different computer but it showed the same result (error), and when I tried to proceed my S2 old files it worked just well using my computer. Could it be my disk space that has a problem or is there any other factor? thankyou
If you have problems with products after the 25th of January (new Processing Baseline 4.0.0) then is can be the memory requirements have increased because of the changed data structure.
Please have a look at the FAQ entry:
I’m getting the error “Cannot construct DataBuffer”, “GC overhead limit exceeded” or “Java Heap Space”. What can I do?
thankyou so much