I am using sentinel-1 GRD products and having following issue:
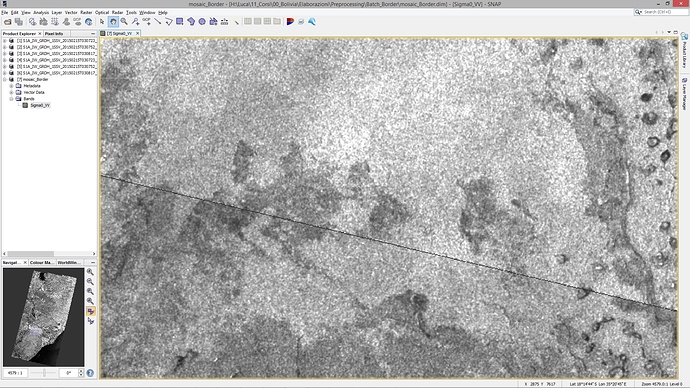
apply orbit file—calibrate—multilooking—subset—terrain correction(range-doppler terrain correction)—SAR mosaic,Everything is ok ,but the mosaic result exists a gap between the two images.
Hi,
I have exactly the same problem by following the same procedure.
is there any solution?
Thanks in advance,
Luca
Can you please provide us with the file names of the products used.
In the past we had similar issue but was fixed, so we need to check if it is similar problem. Thanks
Before calibration apply GRD border noise removal. In the mosaic use
weighted average of overlap.
Also, for performance, it’s better to not include mosaic in your graph.
Do it separate and the performance will be better.
Hi,
first of all thanks for your prompt replies.
I tried also applying GRD border noise removal and follow what Iveci said but I still have the problem.
Michael, the file names are:
S1A_IW_GRDH_1SSV_20150215T030723_20150215T030752_004628_005B47_637E
S1A_IW_GRDH_1SSV_20150215T030752_20150215T030817_004628_005B47_9377
S1A_IW_GRDH_1SSV_20150215T030817_20150215T030842_004628_005B47_C15C
If it may help, I am adding a screenshot to show the problem.
I look forward to hearing from you.
Cheers!
Luca
Hi,
I am using snap 2 beta 8 with the following step
Remove border - cal-multilook 2x2-terrain flattening or not - range doppler terrain correctiin and when i display data without mosaics I have same problem with a gap of 1 or 2 pixel between slice whatever the area among 5 different swath.
Best
Luqa, the products you specified are slices. Use the S1 Slice Assembly operator. I did not get any gaps.
Thank you for you replies. the file names are:
S1A_IW_GRDH_1SDV_20150217T222245_20150217T222310_004669_005C3C_9A27
S1A_IW_GRDH_1SDV_20150217T222310_20150217T222335_004669_005C3C_CD03
Continuing the discussion from SAR mosaic:
Hello,
about the slice, when I apply it to some dataset:
edit:
S1A_IW_GRDH_1SDV_20150503T094559_20150503T094624_005755_00763F_68EE
S1A_IW_GRDH_1SDV_20150503T094534_20150503T094559_005755_00763F_5560
S1A_IW_GRDH_1SDV_20150503T094505_20150503T094534_005755_00763F_1133),
as you can see after terrain flattening and or terrain correction the slic mistakes is enhance: it’s like the slice have done overlap between 2 consecutive data (not all). I don’t remember the path number but I will edit tomorow this message.
Do you now this problem ?
At last, I agree that slice solve the gap problem however the counter part is that we need more ram than when we process each slices individualy… for 4 slices it doesn’t work with 8GO ram but with our new workstation (32GO ram) it take 15GO of ram for slice assembly. It’s nice with the new Workstation but best would be to work on standard computer.
Best
Dear Iveci,
thank you very much for your suggestion.
The S1 Slice Assembly operator should be run before the other steps,right?
(Apply orbit file, Calibrate, Multilooking, Range-Doppler Terrain Correction)
Why the Mosaic tool doesn’t perform properly with the slices?
Thanks,
Luca
Hello,
In a new dataset in Peru with 2 concecutive data, slice on original grd data does not work with the message “you need 2 concecutive data”. However after applyin the slice on the data with precise orbit all was working well.
So, I need to test that in the example where I give the screenshot because may be the Slice problem on some dataset is due to orbit accuracy.
Keep in touch.
Best
Hi,
some news about my slice problem.
The new test was to do the following things based on 3 consecutive data (same slice):
- Apply precise orbit - Remove border - cal
- Next slice assembly
- Next ML 2x2 - terrain flattening - terrain correction
Results…Perfect contrary to begin by slice.
My conclusion, even it’s not totally rigorous is that the problem seems to came from orbit.
On the other side I try to mosaic this data without slice but with precise orbit and I have always the 1 or 2 pixel gap between the data.
Best
Hi,
thanks clardeux! I followed your workflow (though I work with GRDH and don’t apply precise orbit and multilooking).
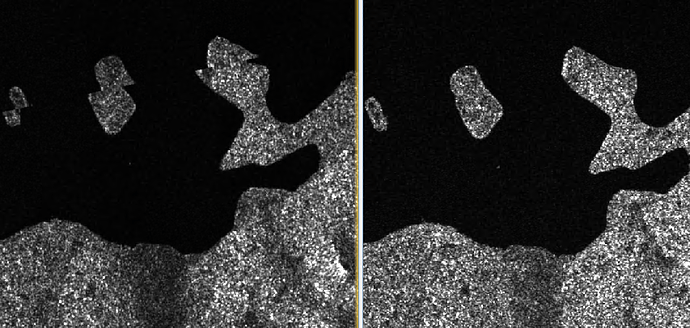
The stripes are gone but unfortunately I have quite unfavorable offsets now.
see screenshot: mosaic on the left with slice assembly and on the right without slice assembly.
Anybody has a good idea how to work around?
best
Esther
OK, again, thanks clardeux, I got it now (I hope):
with S1 GRDH also the orbit needs to be coorected
-> apply orbit file -> S-1 Remove GRD Border Noise -> Radiometric calibration -> S-1 Slice assembly -> Radiometric Terrain Flattening -> Geometric Terrain Correction -> SAR mosaic (with 500 feather pixels and no weighted average of overlap or normalization)
Now the ‘quasi’ horizontal stripes are gone and offsets disappear. I am still fighting with quasi vertical stripes where the mosaic stitches. But will open another question for this.
Best regards
In order to have smooth transition between neighboring orbits you need, either to increase the border margin limit during border noise removal either (alternatively) just to subset/crop the scenes. Then during mosaic there will be no mixing between real data and non-data (black border) in range.
Thank you mfoumelis!
I tried with widening the border margin but still have singular pixels on the left and right boundaries of the scenes.
Now subsetting: I fear the tool is not batchable because I cannot simply say ‘remove 1000 pixels on left and right side’ but have to specify the new image extent for each single scene.
I am currently testing this and will post further If I find out something. But I would still be happy for other ideas.
Best
Esther
Dear all,
I read your discussion and I have same problem.
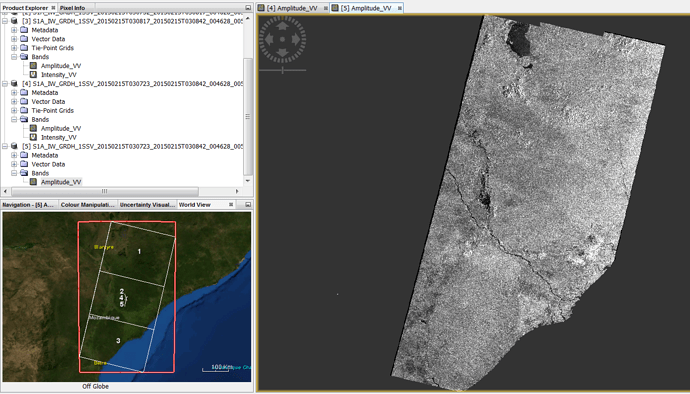
I have tens of consecutive GRDH images in couple of orbits. I use “slice assembly” and “SAR mosaic” at the end. I did not get images in an expected extend, like what Mr @lveci got.
The output dimension is almost half of what it should.
When I add Calibration, Terrain correction, … after ML, still I got same output. The 4 inputs are consecutive image pairs in 2 neighboring Descending orbits, so image should look like square.
example:
Any idea for a proper processing chain to get SAR mosaic?
Is this right? Apply orbit file> Remove GRD border> Slice Assembly> Multi-look >Calibration> Terrain Correction > SAR mosaic
When I apply it, also did not get proper result. Can anyone please help?
Thanks in advance.
I would do slice assembly before anything else as it reconstitutes the satellite data-take.
Hello,
I have a problem doing mosaic using SNAP with S1A & S1B W1_SCL product , I did all the steps
download, regis, forma, beckgeo, debru, remov, filter ,export, unwrap, import andT correction. The fringes are separated,and I would like to join them to see the whole pattern. I did the mosaic but I get black area between the images.
any suggestions please.
Hello,
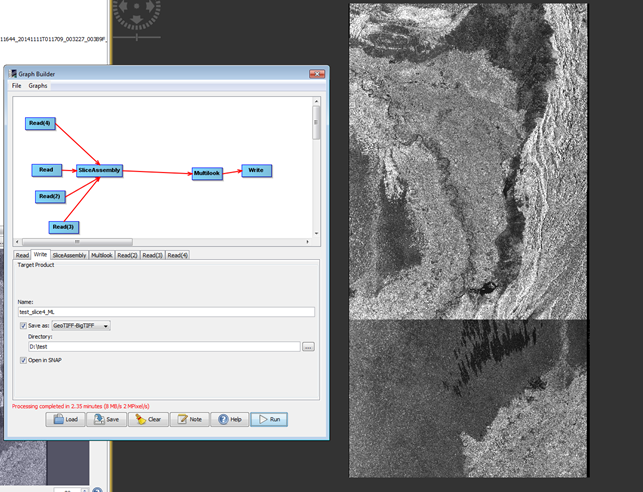
Ive been looking at this issue myself, I have a Graph which currently carries out slice assembly, quite well, but Im running into some issues.
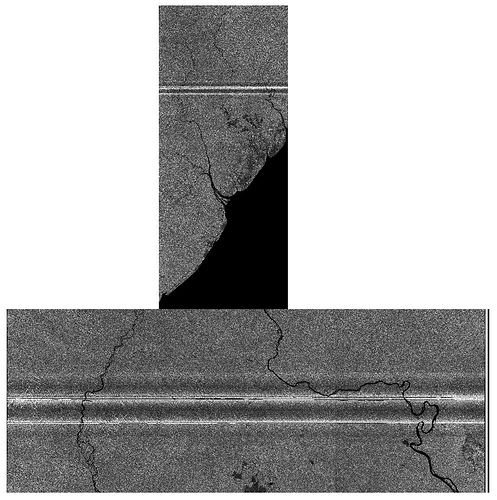
The Image below is an assembled slice based on 3 Sentinel 1 products, I have highlighted the problem area.
My issue is I am trying to Mosaic the two assembled slices, but unfortunately get this as a result
Any help appreciated.