XML is not an image, it is a text-file. The images of Sentinel-2 are *.jp2 files in the folder “GRANULE”
you can export it via File > Export but problable you will have to resample it to a common resolution (10, 20 or 60 m) first.
oh,i got it …
so i have so many questions:
(1) as you see below,when i use the sen2cor to deal with the sentinel-2 images, i choose the resolution 60, did it mean that all the bands’s resolution of the output product is 60???
(2)how could i know the resolution of the output product???
(3)you have said that i should first resampled the output product, and then save it as the tiff type,so how could i resample the image in SNAP???
thank you !!!
thank you !!!
you set a resolution for sen2cor at wich your target product gets written (eg. the scene classification)
Just try to export your desired band and if there is an issue with resampling you get a message wich directly leads you to the resampling dialogue.
you mean that i choose the resolution 60 ,all the bands resolution is 60 meters???
you mean that i choose the resolution 60 ,all the bands resolution is 60 meters???"when i export it as tiff, it shows:::
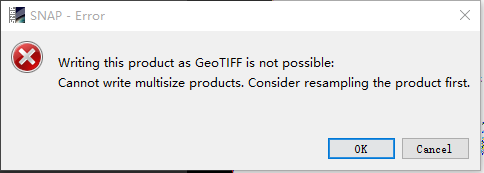
when i click ‘ok’, it did not lead me to resample the image …terrible…
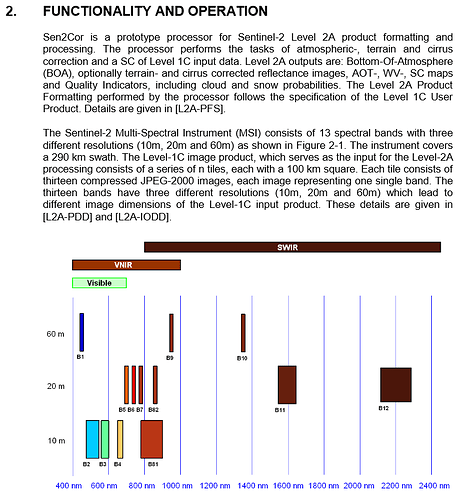
have a look at tha data structure of S2 data:
Source: http://step.esa.int/thirdparties/sen2cor/2.2.1/S2PAD-VEGA-SUM-0001-2.2.pdf
Some bands are natively in 10m/20m, some in 60m. If you want to display the product, all pixels have to be of same size. Therefore, each band exists in all three resolutions.
The size you enter in sen2cor selects which products are used.
However, each band will be created in its native resolution by sen2cor. Meaning that if you want to export your data you will have to resample it to a common resolution byRaster > Geometric Operations > Resampling
Please read the sen2cor manual and use the SNAP help menu.
ok,i will read it carefully
i have a question: i have processed the sentinel-2 images using the sen2cor, and then i resampled it at 60 meters, and save the images as the TIFF,and then import images to ENVI software,we can see as below,it has 34 bands,
and then i count the bands number of the resampled tiff image in the SNAP software in the file name ‘bands’::::
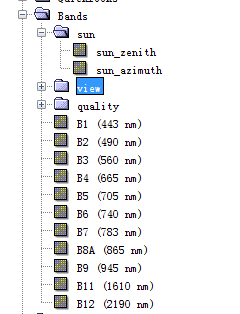
and it also has 34 output under the file ‘band’,so i guess which of them are the bands i need ???
maybe you can create a subset of your data first which only includes the spectral bands you need.
Dear litingting,
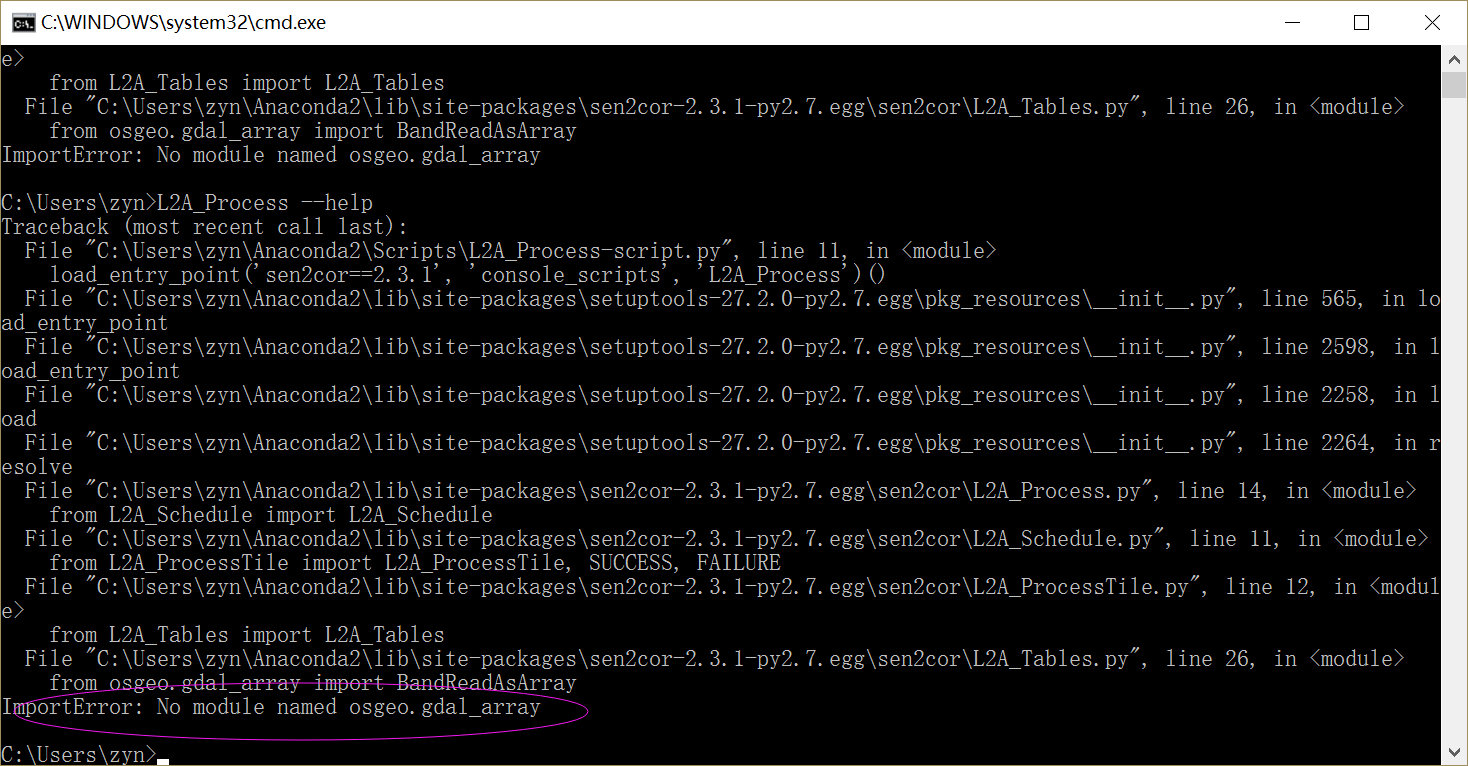
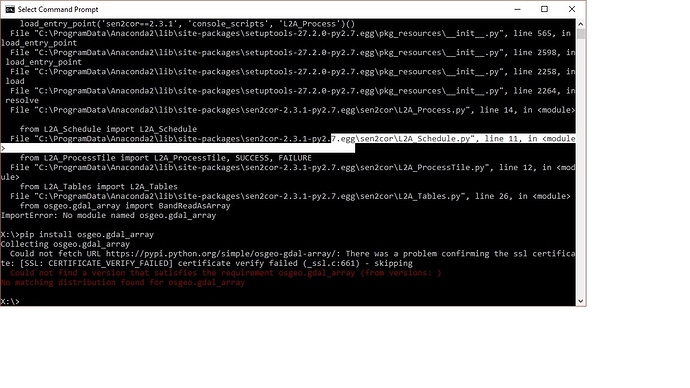
Have you solved the problem?I met the same question when installing the Sen2cor.I installed the Anaconda2-4.3.1,and then I installed the Sen2cor(Version 2.3.1).When the installing process was finished,it printed that‘succeessfully install’.But when I called the processor by"L2A_Process --help" to prove if it really was installed succeessfully ,it printed that“ImportError: No module named osgeo.gdal_array”. I don’t know where the problem is. Any help whill be appriciated.
Yanny
you can type “pip install osgeo.gdal_array” to install this module
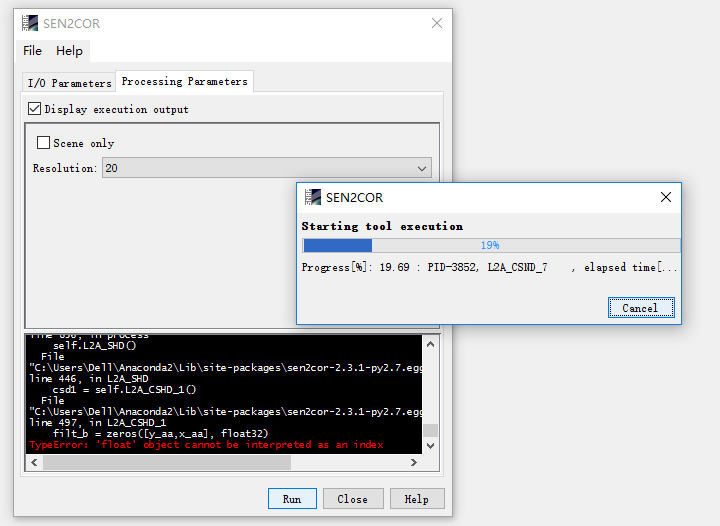
i have installed the latest version of SNAP, snap 5.0 and then i use the software to deal with a 2017 sentinel data , but the error occurs as follows:
what should i do?
@litingting I ‘ve solved it in other way,but thank you all the same.
Can you provide some details? I’m having the same problem, osgeo.gdal_array missing and nothing seems to install it.
I have the same problem without osgeo.gdal_array. I’ve tried to download it from the command line and even checked the url manually, but only found that the page doesn’t exist.
Hi! Have you managed to solve the error “No module named gdal”??