When trying to apply the SEN2COR to a S2 image, the following error appears: "Path does not exist: ‘python.exe’ "
What to do now?
When trying to apply the SEN2COR to a S2 image, the following error appears: "Path does not exist: ‘python.exe’ "
What to do now?
You need to install the SEN2COR software independently from SNAP.
From SNAP Help, follow Sentinel Toolbox Application>Optical>Thematic Land Processing>Sen2Cor Installation.
You will find instructions about where to download the tool
Thanks Julien,
I have followed the steps as given in the PDF, such as downloading Anaconda, unzipping, and then running it into Anaconda environment.
However, when then repeating the SEN2COR tool, a new error message appears. “Error reading product…(and then follows the name of the S2…xml) file”
And then follows:
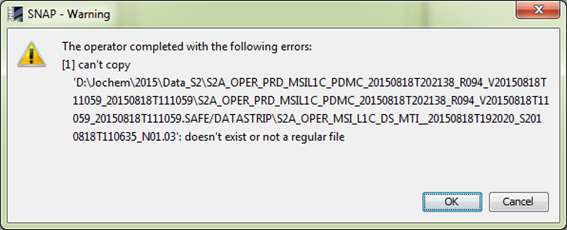
Any ideas what went wrong?
Thanks for help!
I have the same issue with different S2 tiles as well. I also tried the Sen2cor command line version, but received the same error at the end of the stack trace.
I’d like to add that I got an ‘error reading product xxx’ window before the warning above popped up.
I think same problem here for me although strangely if I change the Processing Parameter resolution to 10m or 20m from the original 60m if will provide a different error and then produce some sort of output file with a subset of the original bands - see next post
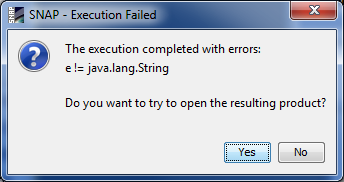
@JochemV & @BBrede: This looks like a Windows path problem; the product has to be in the root directory on Windows < 8, otherwise the path to most files are not valid paths. Actually you won’t even be able to move your product as Windows won’t consider these files as valid.
Dear Nicolas,
Thanks for your reply. Does that mean I cannot store my S2 images where I wish? So if I understand correctly I have to put them in the newly created ‘sen2cor’ folder? Is there any manual that gives all that info?
Thanks.
Jochem
I have experienced same problems.
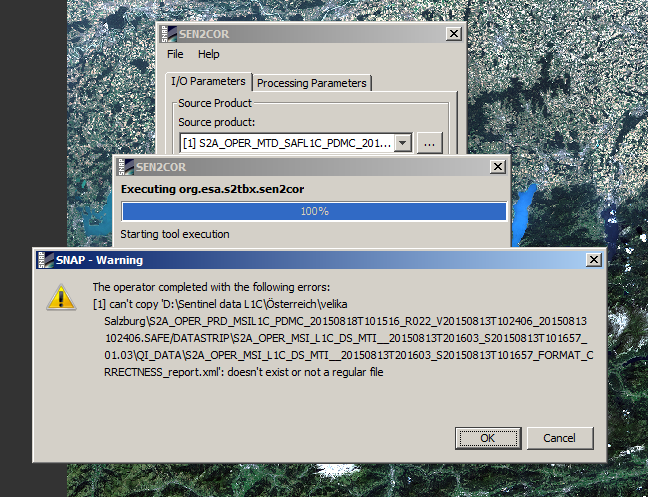
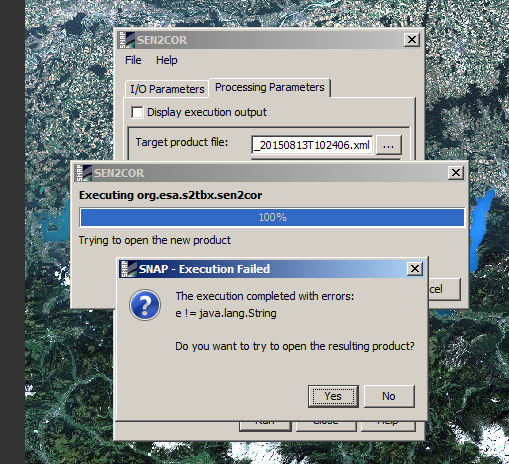
Cheers,
Milos
I am also confused by these so-called Windows path errors. I am used to store data in the D disk, and having the software installed in C.
But @Nicolas seems to suggest that it is not the right way to do?
Some clarification would be welcome.
Kind regards, Jochem
Hi, I’m using the command line interface and get similar error messages:
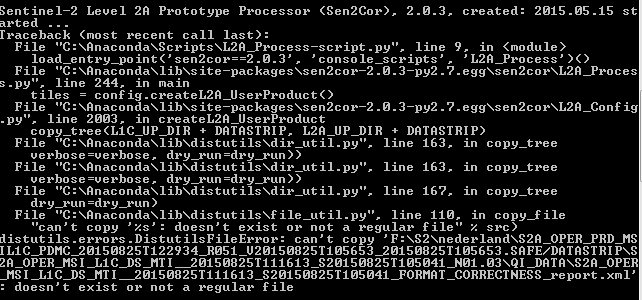
@JochemV C: or D: is fine, but products should be stored on the tom
folder, in D:, not in D:\Jochem\2015\Data_S2, otherwise the full path
to some files is too long on Windows WP/Vista/7 which limits path to 260
characters max.
Thanks @Nicolas. I understand. So I located the output immediately to D. However it caused me running into a new error after the message ‘Starting tool execution’. Unfortunately it does not provide much information. Any idea?
Thanks!
Jochem
Hi all,
The suggested action to place the data folder directly in the root works for me (F:).
Thanks for the suggestion.
I have run the SEN2COR which seems to work and it exits and produces files.
Unfortunately the program does not produce sensible values above surface water (the output bands contain constants above surface water).
Is there some setting that could be changed to force output above surface water?
Thanks,
Steef
Another question.
Is it possible to uninstall Anaconda? That package was required for installing the SEN2COR tool. However, it is about 1.5 GB big. I consider that big, and if no further used, I prefer to have it removed.
kind regards.
Jochem
Hi,
Executing sen2cor from c:\ on a windows 7 enterprise edition 64 bit workstation. Get message Jochem gets that execution failed and then followed by
suggesting file name is still too long. I can adjust file names in the SAFE file but this is tedious since there are many places it would be required. An output file is created but it does not seem to have any Level 2 metadata - just the bands at the resolution I requrested that seem to be same as Level 1C input. Any suggestions please.
Regards
Richard
Hi Richard,
My workaround: I always just renamed the folder of the INPUT image from this really long name to something like “S2A_1C_2_VENICE”. Just remember to keep the “1C” part in the folder name, otherwise SEN2COR is complaining. The ouput image is automatically named in a similar way, so the path is kept shorter and it works without any problems!
BR
mischa
Hi Mischa,
Thanks for your valuable suggestion! I renamed the folder and now for the first time it was executing SEN2COR.  For the developers, it would be extremely useful if you can find a way to cope with with the original image names - each time manually changing the image name is not the most elegant solution. Alternatively, mention it clearly in a manual.
For the developers, it would be extremely useful if you can find a way to cope with with the original image names - each time manually changing the image name is not the most elegant solution. Alternatively, mention it clearly in a manual.
However, because the XML kept the original name, it completed with the following warning without displaying the corrected image:
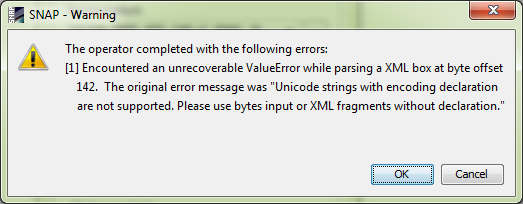
I still did not reach to a corrected image, but it seems we are getting there.
grtz. J.
“I still did not reach to a corrected image …”: its not quite clear what you mean with this last statement. If it is the problem with the mismatch in the color representation (see https://senbox.atlassian.net/browse/SIITBX-113) then
change the DN Value in the L2A_GIPP.xml located in sen2cor/cfg in your user directory to 2000. It is set by default to 4000 as earliest test images did have this scaling. This will probably cure your problem. See my last comments on: https://senbox.atlassian.net/browse/SIITBX-113
Hi Uwe,
It did not reach to a corrected image. Before the error mentioned above, another error appeared, probably related with the long name:
Meanwhile I switched to a laptop with Windows 10. There I also had to shorten the name otherwise leading to the earlier-mentioned error right away. After then running for abouth halve an hour and reaching 100% the same error message appeared (first the Error reading prouct and then the SNAP warning message). So I am still without a corrected image.
Hopefully the SEN2COR tool will be made more user-friendly and accompanied by a manual.
kind regards, Jochem