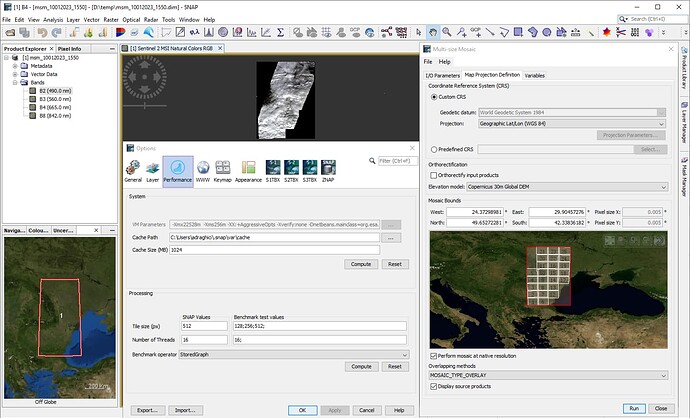
When I process 6 S2 zip files for the neigbouring datastrips of the same orbit with SNAP multisize mosaic processor, it will take extremely long time ,almost take 3 or more days, but similar mosaicking withENVI will take about 30 minutes or less than one hour (tested with 10m bands). In fact , I want to mosaic S2 data for more than 20 consecutive or neighbouring datastrips of the same orbit within 2~4 hours timespan, but it seems SNAP can’t sustain this processing load. Can anyone provide more efficient way or walkround about this requirement? Thanks.
Hi, I’m facing the same problem. Did you find any workaround?
Thanks
No, I haven’t found walkstrough or got tips yet. I hope SNAP development team can have an insight into the implementation of mosaicking processor ,or have a test on 20~30 consecutive images( each is the standard data frame of UTM 100km * 100km) to see how long it will take.
How much memory are you giving SNAP? I suspect that it makes a huge difference in this case.