Dear all,
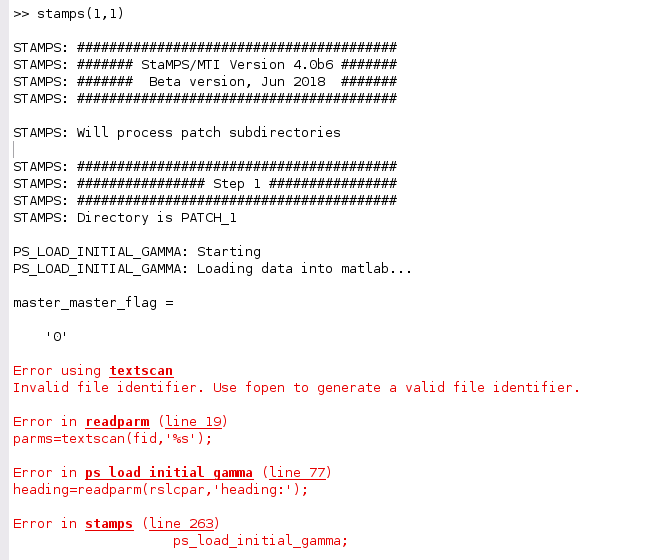
I am running into an issue with 1st step of StaMPS processing - loading data into matlab. I am working with a Ubuntu 16.04 VM created using Virtual box. I have Matlab installed as well as StaMPS 4.1b and all the required tools. I have used the snap2stamps workflow for preprocessing the data using snap v7. All steps so far have run without problems and the interferograms all seem to be correctly created. However when I run the stamps(1,1) I get following error.
I have found a single reference to this error which suggested to use Linux, which I am using. Has anyone encounered this before? I have also reinstalled matlab and tried to see if all StaMPS scripts are the latest version, but without result.
Moreover the code above references version 4.0b, but I ahve never installed this version. I have downloaded and installed this StaMPS on clean VM few days ago using latest version dowloaded from the Github.
Another question I have is regarding preprocessing with SNAP 7. I have been recently told that there are certain “issues” but I have not been able to find a single reference as to what these should be. With the exepion to an update to the ESD parameters. Could anyone provide more information about this, if it is the case? Could the above error be caused by data preparation in SNAP 7?
I would be wery gratefull for any help. Thank you very much.