ok,i will try it
pathlib is part of the python standard library, so it shouldn’t be necessary to install it. Although it was only introduced in Python 3.4. Maybe you are running an older version of Python? Does
python --version
give a version equal or greater to 3.4?
Well, pathlib should be installed when using python 2.7, as it is not in the standard library as it is the case for python 3.4, as mentioned by @gbaier
The current snap2stamps package was developed using python 2.7 and hence, they may exists some issues while trying to run it using python 3.X
If you have anaconda with python 2.7, it includes already pip, so you should install pathlib just by using : pip install pathlib as written in the manual.
If any of you may try with python 3.X, please provide me some feedback to understand which functions may not be compatible, so I can try to use another ones in the future.
according to my experience with Anaconda (python 2.7) and these pages, all Anaconda distributions include pathlib:
https://docs.anaconda.com/anaconda/packages/pkg-docs/
You both may be right! Indeed I have just checked that I installed the required library on the system, and not in anaconda. Sorry for the confusion!
Thanks again for keeping it sharp! 
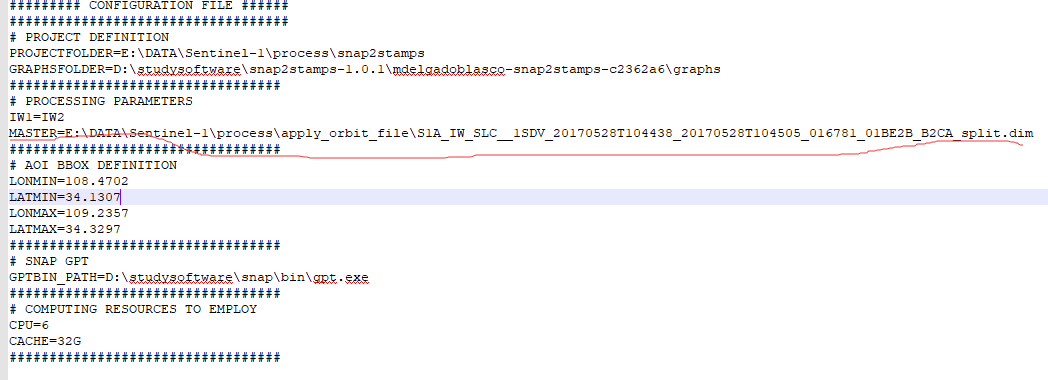
two question when i process 30 images(1 master+29 slaves) by snap2stamps tools the project.conf file i used is like this:
the master file is the one that had been splitted and applied orbit file in snap, is this exist any errors?
and i finished cored_ifg_topsar.py
the config file looks fine. Just make sure you call the scripts in the correct order
- prepare slaves
- split slaves
- coregistration and interferogram generation
- export
ok,a question about para settings of snap disurbes me in the manual:
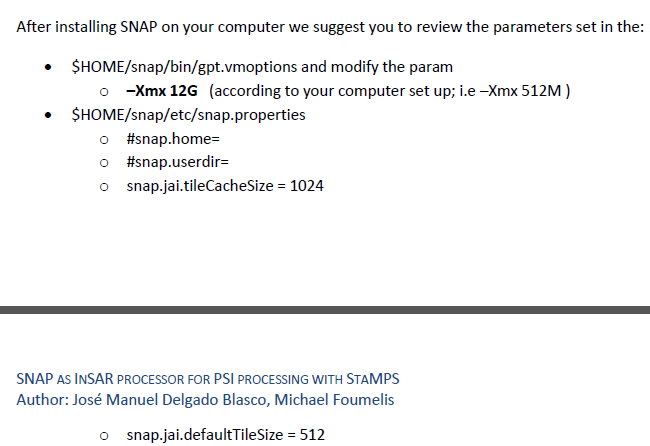

in my gpt.vmoptions file,I set the XMX to32G as my computer memory space,and i dont konw konw how to set the tileCachaeSize and defaultTileSize acoording my compute
Dear @zhuhaxixiong,
I also have 32GB RAM computer and I normally set the tileCacheSize to 8192 and defaultTileSize to 2048.
I hope this helps.
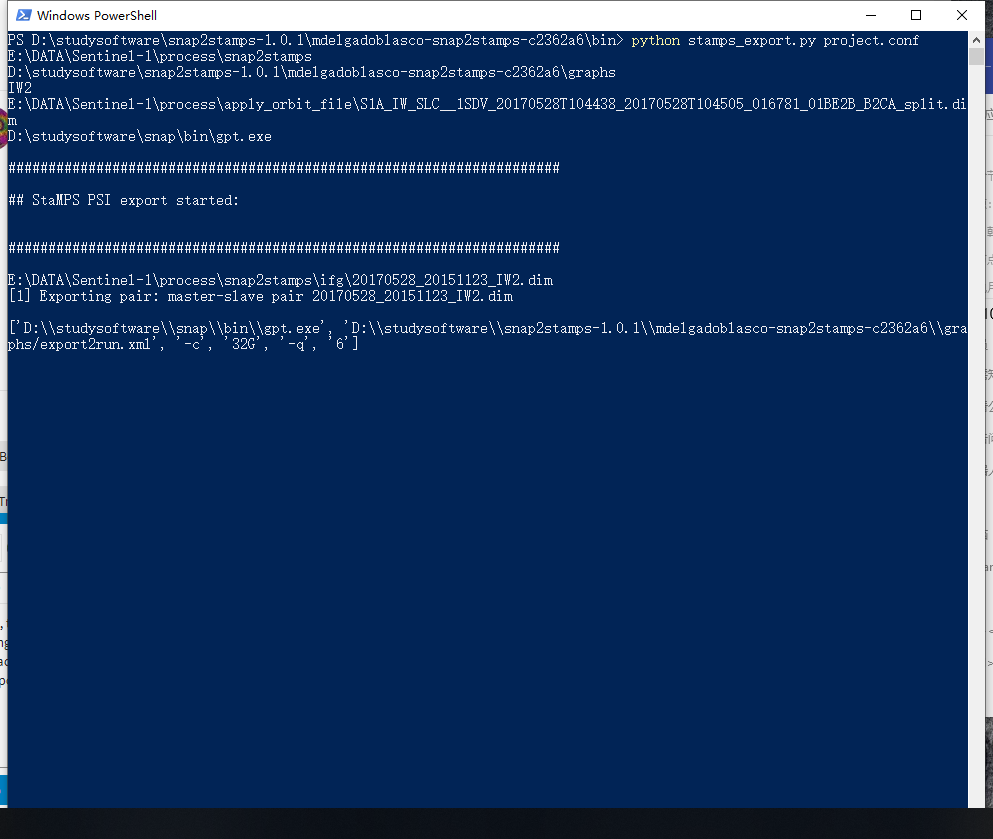
ok, thanks a lot.when i do the final step stamps_export, the process is really long, my friend close my process window by acccident.when i restart my machine and check the file it export, there are only ssome part of ,so should i export again,and the process restart from the first file.
Dear @zhuhaxixiong,
In fact this is a good point of discussion.
It can be included for next versions to check if the file exists or not before process it again, and it can be included as an option that the final user should decide whether to activate or not.
Essentially the main issue to apply or not such check to avoid re-processing again files, we cannot ensure the file found is fully correct or for example was stopped while being processed. If it would be the latter case, the process would start on the next file, not reprocessing the file which processing was accidentally stopped. I am not I am clear enough with it.
In any case, I will include the overwrite option, in which will be the final user the one responsible to check the integrity of the files. For the future I may think a better option to do so. Again, any feedback is welcome.
Cheers
thank you very much
when i use this tools to export ,what should i change in the mt_prep_gamma and ps_load_initial file
My suggestion is to download the current StaMPS version, for which you should change nothing else.
Get it from https://github.com/dbekaert/stamps by cloning or downloading the current version, and not downloading from the release section. Doing so you get the latest updates which help you to deal and avoid potential errors in successive StaMPS steps.
Cheers
and you means i do not need to replace the two files i get from the forum?
In the github package you have everything you need.
In the forum you may find several versions of those files, but within the github package you get them updated.
I hope this helps
ok,maybe there should a new text like"PS Using Manual_3.1" more exactly
Sorry but I dot not get it.
New StaMPS version is 4.1b and it seems well organised to me, including the User Manual.
Enjoy!
thanks alot
Thank you for the work on this script!
I was just going to write my own one from scratch but this is so much better.
I think in the future I would like to remove the .xml graphs and instead just use python + snappy scripts to make it nice and smooth.
Can you please help with the question in the end of this long post?
I have successfully ran snap2stamps in a ubuntu docker container on a server as follows…
First the master output naming convention caught me out.
For now you need to run it in GUI (split + orbit) with more than one burst and name the output file the same as the S1 product with a different suffix.
I think you should just make it part of processing.
Attached is an .xml file you could add, it takes a WKT polygon as input so it will extract only the bursts that overlap the polygon. master-split.xml (1.0 KB).
Run this with an input master scene and a wkt text file as follows (cat just reads the wkt file which looks like: POLYGON((18.16 -32.88,18.53 -32.88,18.53 -33.11,18.16 -33.11,18.16 -32.88)) get it from wkt online)
gpt ./master-split.xml -t “./S1A_XXXX_orb” -PsourceFile=“./S1A_XXXX.zip” -Pwkt=“$(cat ./aoi.wkt)”
I run the entire snap2stamps with a bash script wrapper as follows - I think snap2stamps should also do this. It can source the wkt from the project.conf file.
#! /bin/bash
gpt /home/scripts/master-split.xml -t gpt ./master-split.xml -t “./S1A_XXXX_orb” -PsourceFile=“./S1A_XXXX.zip” -Pwkt=“$(cat ./aoi.wkt)”
python ./slaves_prep.py ./project.conf
python ./splitting_slaves.py /.project.conf
python ./coreg_ifg_topsar.py /.project.conf
python ./stamps_export.py /.project.conf
I run all of this with a snap dockerfile that I pulled with:
FROM mrmoor/esa-snap-with-python:latest
I also had to install pathlib in the dockerfile with:
RUN apt-get update && apt-get install -y git python-pathlib
I also pull snap2stamps with:
RUN mkdir -p /home/software/ && cd /home/software/ &&
git clone GitHub - mdelgadoblasco/snap2stamps: Using SNAP as InSAR processor for StaMPS &&
chmod +x /home/software/*
In the docker I then run an entrypoint script with this configure snappy:
echo “-Xmx64G” > /usr/local/snap/bin/gpt.vmoptions
echo “snap.parallelism = 16” >> /usr/local/snap/etc/snap.properties
/usr/local/snap/bin/snappy-conf /usr/bin/python
/usr/bin/python /root/.snap/snap-python/snappy/setup.py install
My problem is with the mt_prep_snap script. I use the one that comes with Stamps.
I am attempting to run Stamps with Octave instead of Matlab. Wouldn’t this be amazing?
So I am using Octave v4.4.1 in a docker with the csh shell and I change this line in mt_prep_snap from
matlab -nojvm -nosplash -nodisplay < $STAMPS/matlab/ps_parms_in_itial.m > ps_parms_initial.log
to
octave < $STAMPS/matlab/ps_parms_in_itial.m > ps_parms_initial.log
I get the following output:
mt_prep_snap 20170730 /home/in/131/INSAR_20170730 0.4
mt_prep_snap Andy Hooper, August 2017
Amplitude Dispersion Threshold: 0.4
Processing 1 patch(es) in range and 1 in azimuthoctave: X11 DISPLAY environment variable not set
octave: disabling GUI features
opening /home/in/131/INSAR_20170730/rslc/20150331.rslc…
Segmentation fault (core dumped)
9162
1472
mt_extract_cands Andy Hooper, Jan 2007Patch: PATCH_1
selpsc_patch /home/in/131/INSAR_20170730/selpsc.in patch.in pscands.1.ij pscands.1.da mean_amp.flt f 1
file name for zero amplitude PS: pscands.1.ij0
dispersion threshold = 0.4
width = 9162
number of amplitude files = 0
Segmentation fault (core dumped)psclonlat /home/in/131/INSAR_20170730/psclonlat.in pscands.1.ij pscands.1.ll
opening pscands.1.ij…
Error opening file pscands.1.ijpscdem /home/in/131/INSAR_20170730/pscdem.in pscands.1.ij pscands.1.hgt
opening pscands.1.ij…
pscdem: Error opening file pscands.1.ijpscphase /home/in/131/INSAR_20170730/pscphase.in pscands.1.ij pscands.1.ph
opening pscands.1.ij…
Error opening file pscands.1.ij
So the main error is the Segmentation fault when opening the first scene. The scene looks fine. Wonder if anyone has any ideas? Is this more a question for MAINSAR?
In terms of snap2stamps. Why does it create target.dim and target.data files in the PROJECT directory? I see the data are interferograms, is this then the output before export to Stamps? It would be nice if this had a better filename and output directory.