Hi there,
I am working on a project which needs to retrieve LAI for a huge area (CONUS US). As you may know, it is quite time-consuming to use the biophysical processor. Then, I am wondering is it possible to subset the raw imagery by points since only a part of critical points make sense for me, then do the retrieve. I have tried to use land/sea mask, but it does not work. Does anybody have any experience or suggestions on this? I really appreciate your help.
Yes, this is possible.
In this post some information is collected:
Thanks for your reply @marpet!
I notice that the subset is always mentioned with polygon, but I only need points-based pixels. I currently have a mosaicing image for the whole US. Basically, I want to extract the pixel involved in my research based on a point.shp vector to reduce pixels processed by the biophysical processor. I tried to use land/sea mask, the point.shp could overlap with raw raster but there is nothing/no data in the results raster. Do you have any idea on this?
Ah, okay.
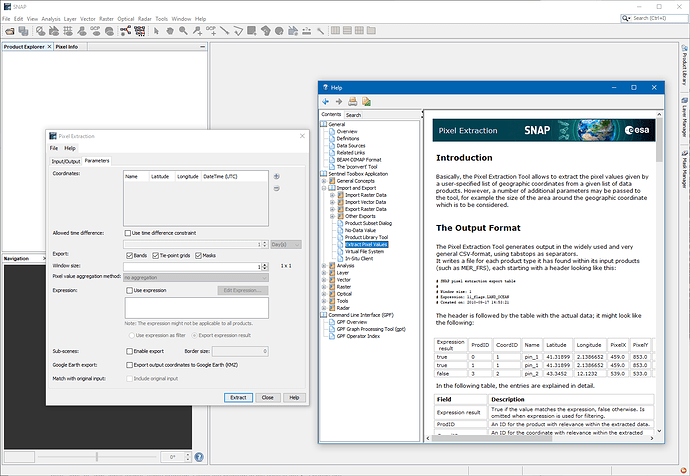
In this case the PixEx tool might be the tool for you.
It extracts the information at provided geo-locations.
You can find it at Raster / Export / Extract Pixel Values
Thank you very much!
I will give a try on this way.
Hi, @marpet
I have tested in a small piece area in this way, and finally, I get a .txt file that contains all bands information. But I am not sure how to process it to run in the biophysical processor since it seems its format can’t be accepted. Then I try to get subscene from “extract pixel value”, but the result image is almost the same as the raw image.
The result can’t be feed into another operator.
But you can use it as last step in a graph. Before you use the biophysical operator.
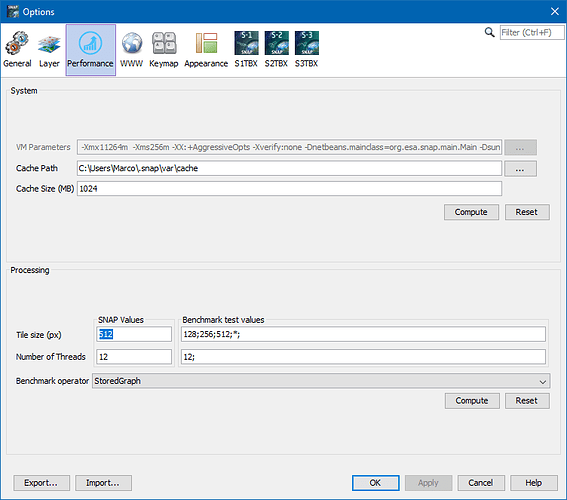
Then only the pixels are computed which are extracted. To be more precise the tiles where the pixels are located are processed. A tile is the smallest processing unit. The size depends on the input format and the configuration.
You can try to configure a small tile size.
But I would not go below 64 pixels.
Bringing the pixels in an order can improve the performance.
The order should be in reading direction. From left to right and then down.
You now that you can compute LAI also by simple ratios
(IDB - Application: Vegetation LAI (indexdatabase.de))?
Sure, this is not the scientific approach but might be sufficient for your use case.
And the calculation is much faster.
Thank you @marpet!
Since I might still need other traits, it is better to use the biophysical processor. I have mosaiced the small tiles (huge number) into a big tile, so it seems that this way doesn’t contribute to it. I will keep trying to look for a less time-consuming way to solve it.