Hi!
I did supervised classification also. To explain what I did, I write down step by step because something went wrong and the result were not what I expected:
- resampling image
- subseting image
- defining geometries
- saving
- reprojecting
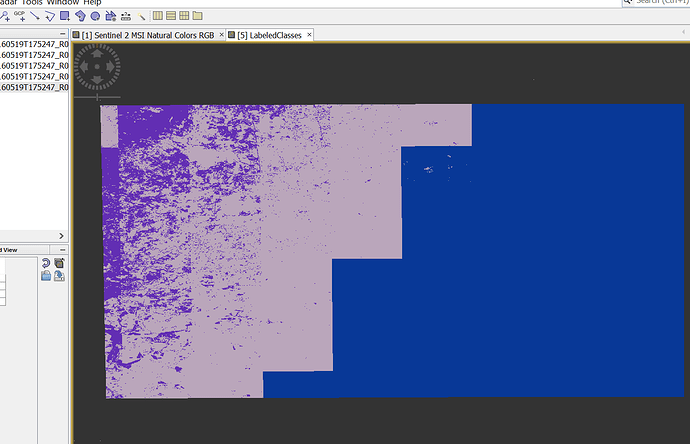
- supervised classification ( KNN-classifier, took 4 hours)
and result
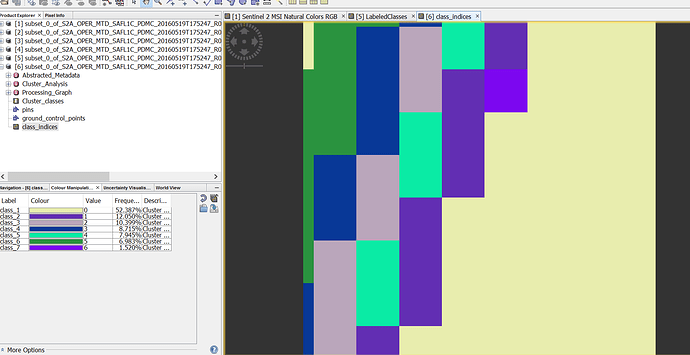
Almost the same (even worse) happened when I tried unsupervised classification ( K-means ):
Any ideas what I did wrong?