I cannot understand this. I tried it myself with the following steps and it works:
@PiriReis: If you follow these steps, it works.
-
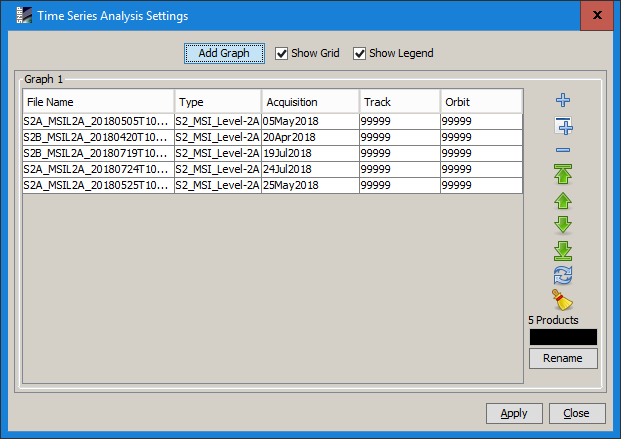
Open 5 S2 images in SNAP
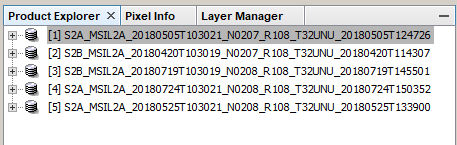
-
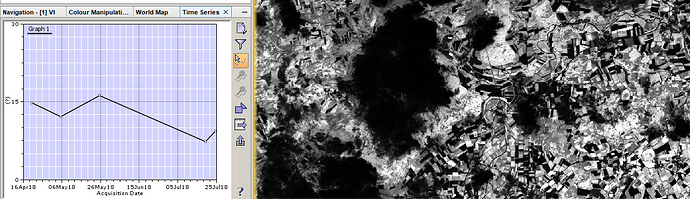
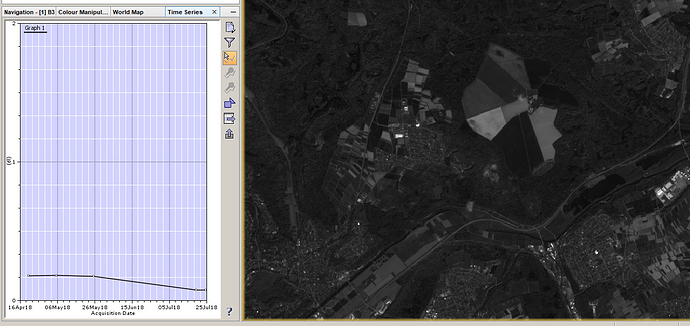
Test the time-series tool for band 3
-
it works
-
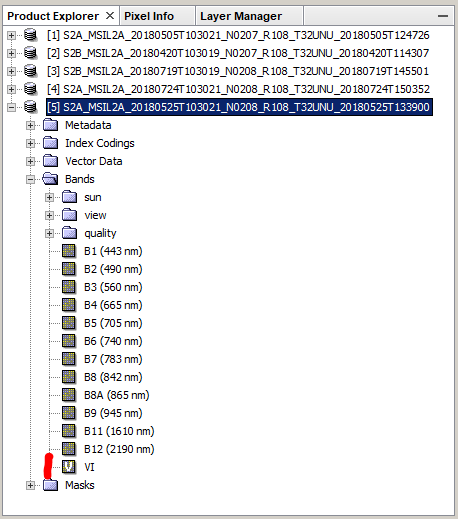
Calculate index (I used the VI [NIR/Red] for the sake of simplicity) for each band
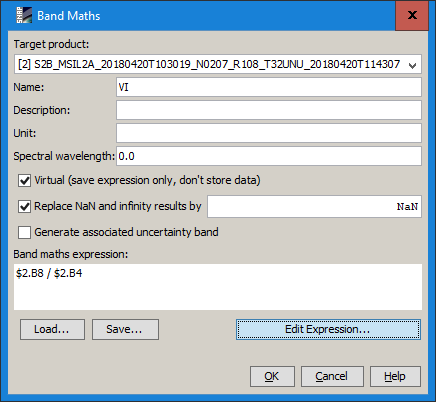
-
band in stack
-
Right-click on new band and click ‘convert band’ so it becomes permanent
-
Save all products (this takes time because the whole products are written into a DIMAP format)
-
Close all products and open again. Clear the time-series tool and insert the products again. The dates are still there.
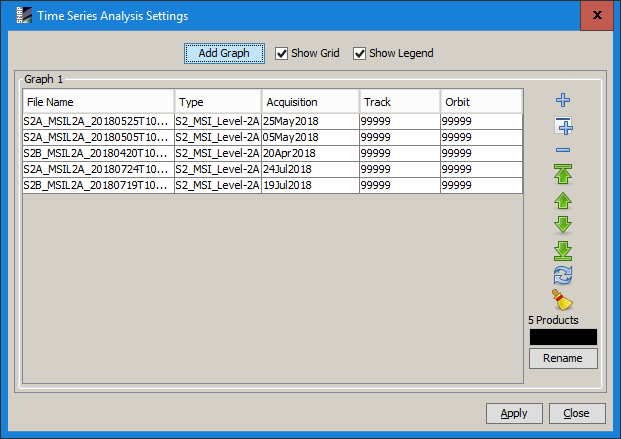 ’
’ -
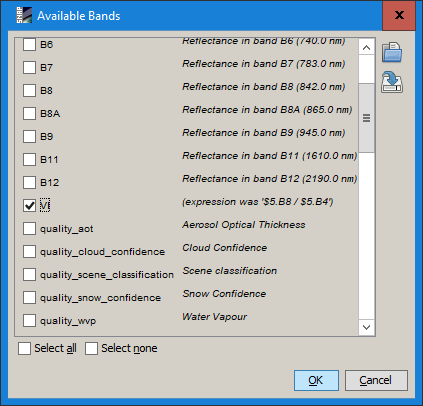
Select the VI in the filter tab
-
Display VI series in the plot (note the tab says “VI” and the scale ranges from 0-30)