Hi @ABraun, is it already StaMPS for R ?? Can you please provide a link for that? Thanks
sorry - fundamental typo: I meant ‘there is no R version of it’
Plz guide how to create stack?
like Master-slave1, Master-slave2, Master-slave3 and so on…
or
Master1-slave1, master2-slave2, Master3-slace3, and so on… i
if feel coherence in 2nd one will be better.
currently, SNAP only supports version 1: master1-slave1, master1-slave-2…
I applied batch processing up to backgeocoding keeping one master suggested by SNAP, keeping master at the top but it has changed master during processing.
Please have a look at this overview: About the STaMPS category
It lists many examples on how people proceeded and retrieved good results.
i fear coherence will lost due to large temporal difference?
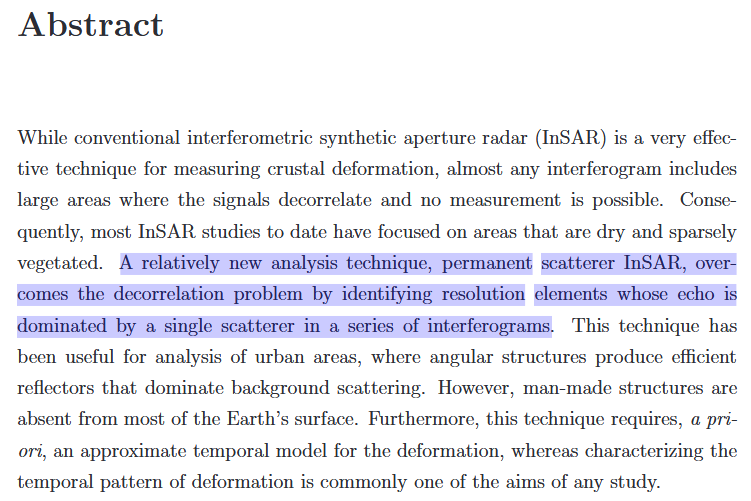
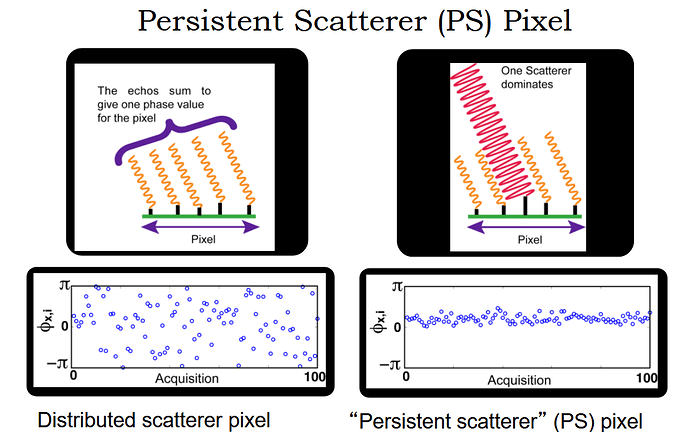
surely, but that is one main advantage of PS InSAR: It is no longer dependent on coherence because persistent scatterers are selected by other criteria:
The PhH thesis of Andy Hooper (main developer of StaMPS) starts like this:
One of his slides:
Source
Hello mdelgado,
I want PSI and have exported a stack of 32 images through SNAP-StaMPS Export. Now the problem is MATLAB is not installed on my VM. I have these resources only.
I’m afraid you cannot run the StaMPS scripts without Matlab, sorry. We told you this in the beginning.
Many academic institutions have license agreements however.
I am arranging a linux workstation for MATLAB and StaMPS.
- Will it work if i download StaMPS exported data only in new system?(rslc, diff0, geo, dem only)
- Do i need to give commands as suggested in chapter 6 of stamps manual? or it will run independently as per [ps_load_initial_gamma(changed).m] and [mt_prep_gamma_snap(changed)] only.
very good 
You can use the data prepared in SNAP for further use in Linux.
First, you call mt_prep_gamma_snap pointing to the exported files to generate the data to be analyzed in Matlab. Then you start matlab and work with the scripts provided in the StaMPS distribution. But in Matlab you only call the different steps [e.g. stamps(1,1)] or change some of the parameters. These are pretty much all the commands you have to give.
Hi katherine
Very thank you for summarizing about using SNAP to Stamps
I do steps 1-8(1-Apply_Orbit_File, 2-stack using Backgeocoding by 8 images, 3- Deburst , 4- Making Interferograms, 5- TopoPhaseRemoval , 6-Add elevation band to only Interferograms bands, 7- DidTerrain Correction (TC) of the two products: master_Stack_Deb and master_Stack_Deb_ifg¬_dinsar
Please see below Figs:
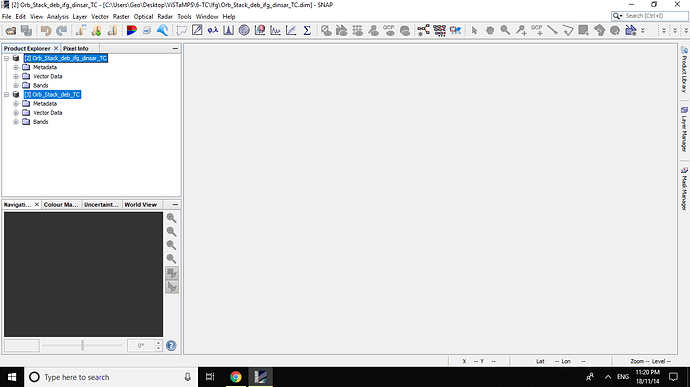
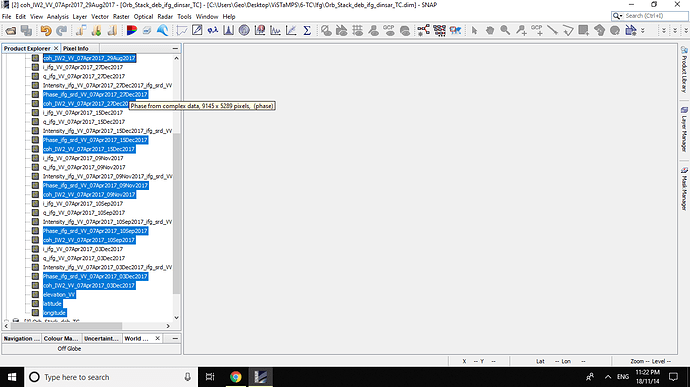
All Bands OF Orb_Stack_deb_ifg_dinsar_TC are :
and All bands of Orb_Stack_deb_TC are
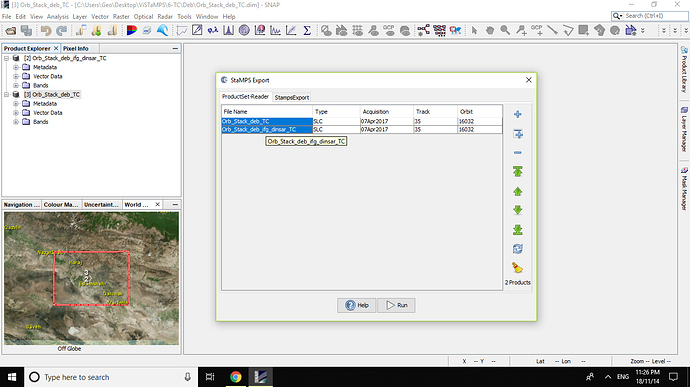
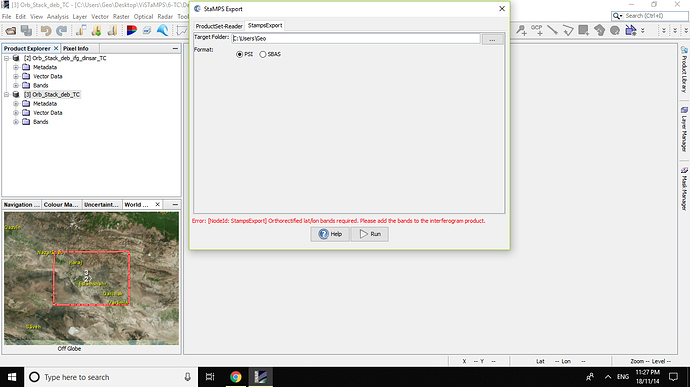
But when i export data for StaMPS , 1 was faced with below erroe and problem:
and
Error : [ Node Id: StampsExport] Orthorectified lat/lon bamds required. Please add the bands to the interferograms product .
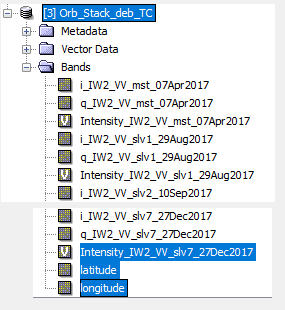
But , as you see, i add lat/lon to interferogerams bands and elevation to Stack Deb bands .
I could not export master_Stack_Deb_ifg_dinsar_TC(with lat/lon and elevation) and master_Stack_Deb_TC(with lat/lon)
Why?
Please help me
Thanks in advance
manually creating the lat/lon and dem bands is outdated, you should select the checkboxes in the topographic phase removal step.
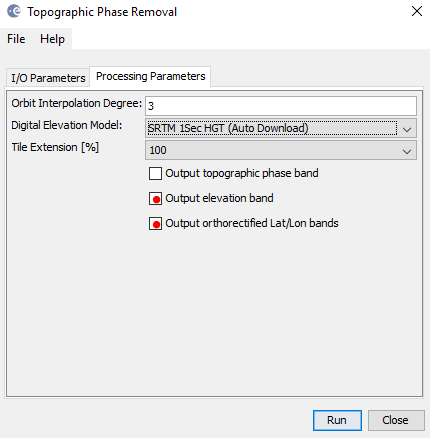
Thank you very much for your quick response
But I’ve already removed the TopoPhaseRemoval in step 5 Without select the checkboxes output Orthorectified lat\lon bands!!
Should I do this again
Is there another way?
very thanks
I don’t know if there is a way around but at least it works with the mentioned checkboxes.
The steps described by katherine were written when these were not yet included in the Topographic Phase Removal module.
thanks ABraun:

Again, it is not recommended to do Terrain Correction before StaMPS.
StaMPS handle the full resolution SLC data, and it is recommended to use the full resolution for PSI.
The TerrainCorrection operator probably it was used in the past when lat/lon orthorectified bands were not produced using SNAP, but again, this is not the case any more.
Sorry for that!
hi
where you ran mt_prep_gamma?
thanks
when you call mt_prep_gamma_snap, you point to the directory of your data which was exported by snap, so it doesn’t matter where you run the script.
At best you create a new folder for the processing and navigate there. All files produced by the script are created here and it is good to have them separately to nothing gets mixed up with other data.