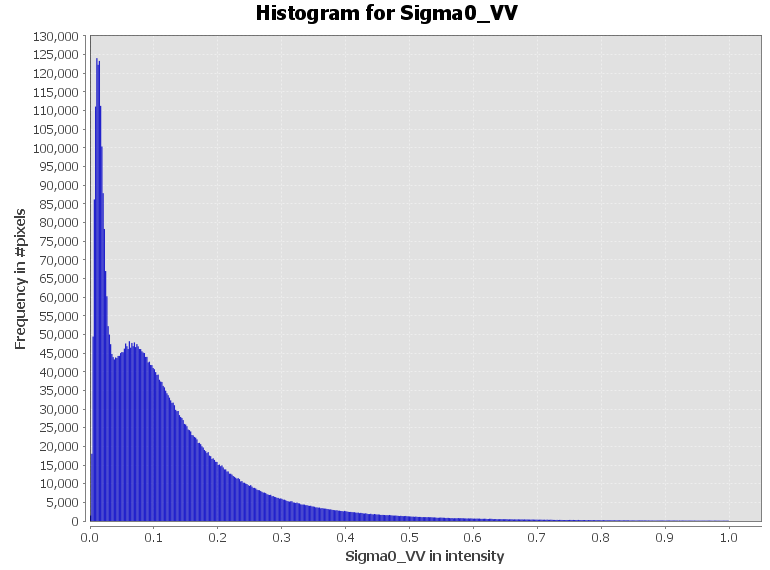
Radar data has a very imbalanced histogram (>95% of all values are smaller than 1, but the remaining 5 % can be impractically large). I would recommend to convert your data to log-scale: right-click > Linear to/from dB
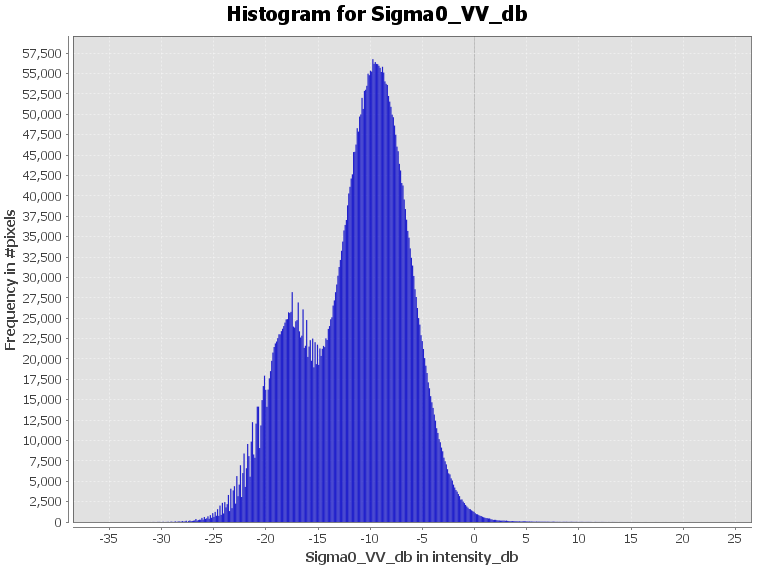
This converts your data to a range between around -40 and +5 with a gaussian distribution of values which is much more suitable for MD or ML classifiers. Additionally, it increases the contrast between black water and bright land pixels.
Use the dB bands as an input for classification and compare if the noise in the result has decreased. If the contrasts are very good, you might even try an unsupervised classification with 2 output classes.
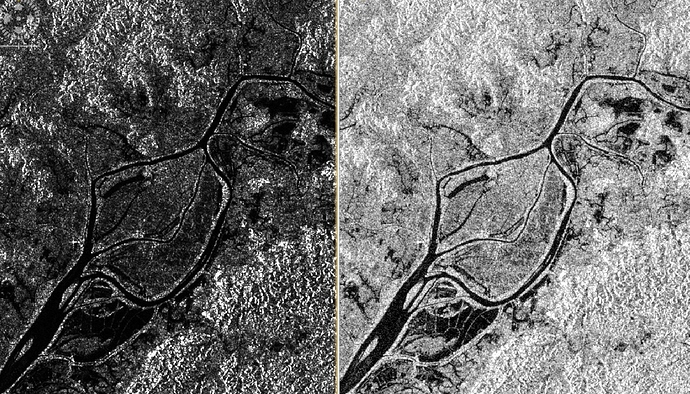
On the left you see the original calibrated data, on the right after conversion to dB scale:
You can clearly see the differences in the histogram: The threshold between dark water and bright land is more pronounced in the dB image.
Maximum Likelihood and Minimum Distance try to find thresholds in a multi-dimensional feature space. So if this feature space is imbalanced, as it is the case for SAR data, they might not work very well.
Besides that, a strong filter (maybe median 7x7) might also reduce the noise over water areas, especially if there are bright pixels because of rough surfaces or partially vegetated water areas.