Hi everyone,
Thanks for helping me, I managed to run the LAI in the end…
but, anyone knows any article about LAI performed by SNAP?
Here is an example of articles concerning LAI
Hi
l have same problem. as l see this data get from Copernicus Open Access Hub.
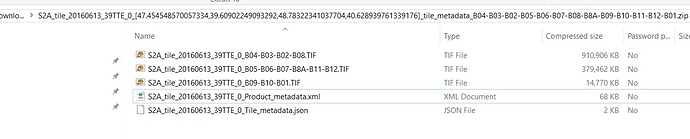
How is about data download from Land Viewer EOS. that does not give MTD data.
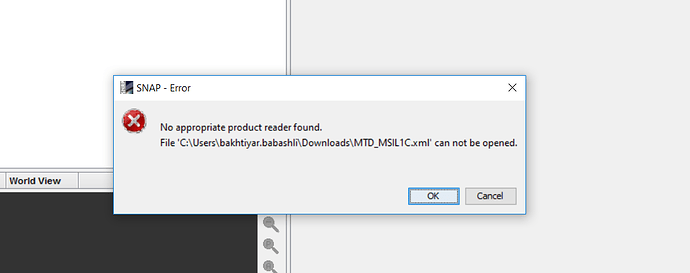
It is provide S2A_tile_20160613_39TTE_0_Product_metadata.xml and this does not opened by SNAP.
what can do in this case?
SNAP needs the S2 data in its original form, e.g. downloaded from the Copernicus Hub, PEPS or other sources which provide the initial product.
The EOS Land Viewer does not offer that (according to my knowledge and the documentation).
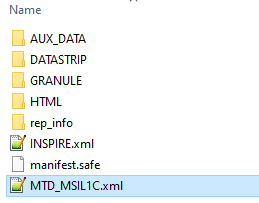
the MTD must be inside the original product folder, e.g.
S2A_MSIL1C_20190904T102021_N0208_R065_T32UMU_20190904T123501.SAFE
which also contains the folders AUX_DATA, DATASTRIP, GRANULE, ect…
this is not a L1C Sentinel-2 product. It should look like this if you want to work with it in SNAP:
Hi,
I met the similar problem when I computed the fAPAR by the biophysical processor. It works well with L1C data. However, when I use the L2A images (.TIF) download from the Google earth engine, it always ends up with the error “Missing band at 560 nm”. But I checked the image and it has band at 560 nm. So do you know what s wrong here? Does biophysical processor only work on 1LC images?
Thanks
I don’t know the GEE data, but if it is tiff the wavelength is probably not specified. Because GeoTiff has no format specification for doing it.
In general the processor should also work on L2A data. But it is best if you get the data from the official ESA sources.
You could add the wavelength information manually to your data.
Right click on the band and select properties. There you will see a field where you can enter the wavelength. Do it for all spectral bands.
ok. I ll try. Thank you so much.