Concerning python any other version rather than one of both Python 2.7 and Python 3.0 (conda), dosen’t help,
is this regarding the installation tips in the git?
i needed 3 and 2 both, for snap2stamps and data downloading
I’m asking generally, if the advance versions of python 3.5, 3.7 are compatible with SNAP, (In case both are important) would you please to explain how to get two pythons in the same machine?
I would not know whether python 3.5 or python 3.7 show problems with the command interface that snap2stamps uses of SNAP, i dont think so.
getting 2 versions of python on the same machine can be done using Anaconda. you can create environments on your pc which is basically multiple python installations in a seperate folder with specifically the packages and versions you need. this way i can hand over my version file to you,you can install a conda environment based on that version/package file and there will be no problems running my scripts.
check anaconda documentation if you want to know more about it.
That’s would be great,
i am sorry, i think you misinterpret my intentions. this is just an example of what is possible with conda environments.
you can download and install miniconda, add conda.exe to your path variable and just run:
conda create C:/users/home/pyth_env2.7 python=2.7 package1 package2 package3 etc
That’s mean all the versions of python are available within miniconda and by adding the conda.exe to my path, all latest versions could be run?
by downloading miniconda you can indeed, MAKE a pyhon environment for all versions and with all packages you want. you can also add packages to the environments if they are not complete anymore or when you need extra things. you can then use the python.exe files in command prompt or open the complete pyth_env folder in an interpreter like Pycharm to make scripts and run them using the interpreter.
Would you please to clarify this point, in the following says don’t check it
Source : https://realpython.com/python-windows-machine-learning-setup/
However, in the following video, he prefers to check it,
So which one to follow?
Also to add the python to the SNAP during the SNAP installation, which one will be the path to python,
I have never added my path to Python in SNAP at the installation but just using the SNAP command line and some steps i found online, and even then i stil have not used python inside SNAP so have no clue.
Regarding the installation of anaconda: you can always add anaconda to your path env variable later if you want to, only differences is that to run conda you will have to type the full path to the .exe file, or be in that directory instead of just typing conda in any directory. i thinki have not registered anaconda as my standard python3.7 but i never use just standard python so honestly i would not know. when doing anything with python i use a specific environment created for it.
I think it’s important to add up the path, because it’s needed to run from cm, some python scripts, for easy example I download images using *py
Please install csh interpreter
Regarding the SEVERE message from the path… you can export the path on the enviromental variable including the . into the LIBRARY_PATH variable. Still this does not produce any error while running the snap2stamps scripts.
I hope his helps
I’d prefer the first one “Add Anaconda to my PATH variable” because it makes python scripts executable from any location.
The second one can be critical if you don’t fully control the versions on your PC. For instance, ArcGIS has its own Python, so does ERDAS, QGIS and so on. If you make the Anaconda environment the absolute first, the installation of packages can ruin pre-configured environments of these programs.
You mean to check the box, as it explains in the video, because the first option says uncheck the box “add anaconda…”
the first one gives you less trouble, the second one needs more control - just my personal opinion.
Hello dear members of this forum. I want to thank to all persons who help to each other.
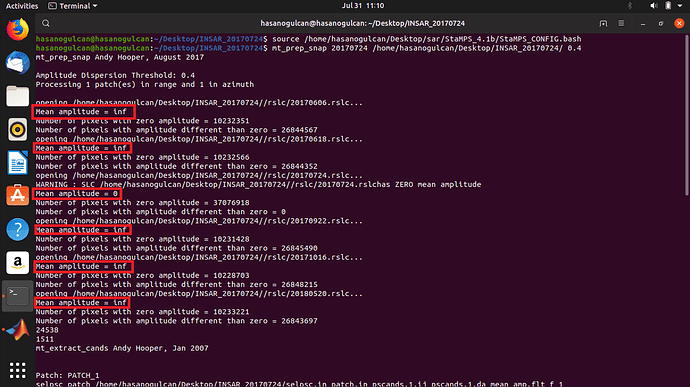
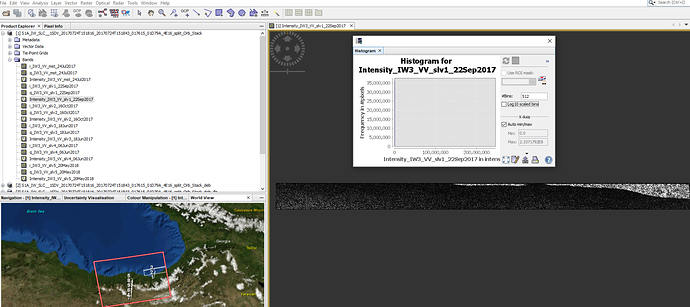
My problem is picture install below. i m working with latest version of StaMPS under linux 19.04 system and SNAP 6.0 . Initially only for trying the PSI process i chose 6 S1A-SLC images totally (1 master+ 5 slaves). Unfortunatelly getting an amplitude value like ‘inf’ and ‘0’. I think this is a problem and trigger the matlab proccesses as negatively. [I would like to set time series and see the deformation on an area on the floor by years]. Thank you. Best regards… @mdelgado, @marcu, @ABraun, @gabrielaquintana77, @FeiLiu, @katherine, @bayzidul
For what you show here my guess is that the coregistration resulted in corrupted data (NaNs) and for this reason then you get Inf as average amplitude.
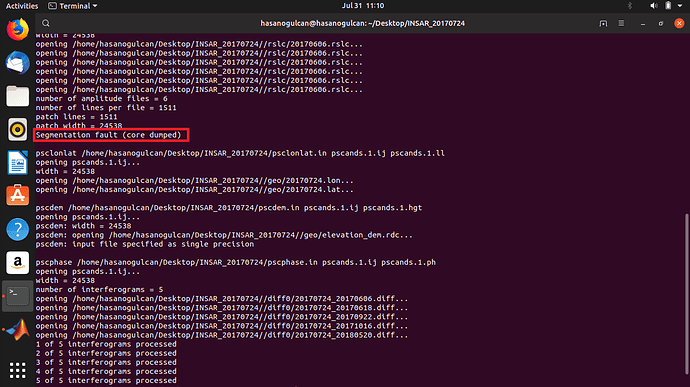
The second issue of Segmentation fault should be linked to RAM available in the computer.
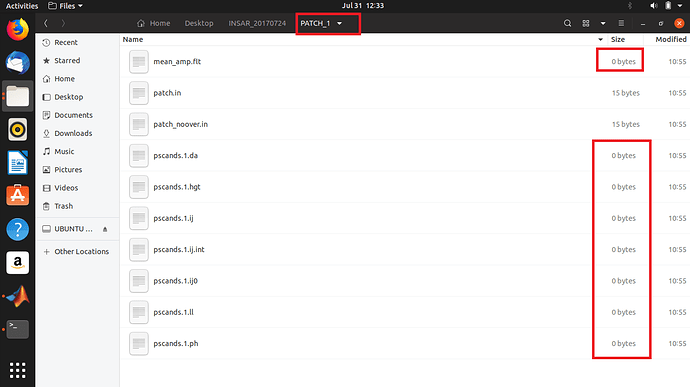
Obviously the 0 bytes are linked to all previous issues.
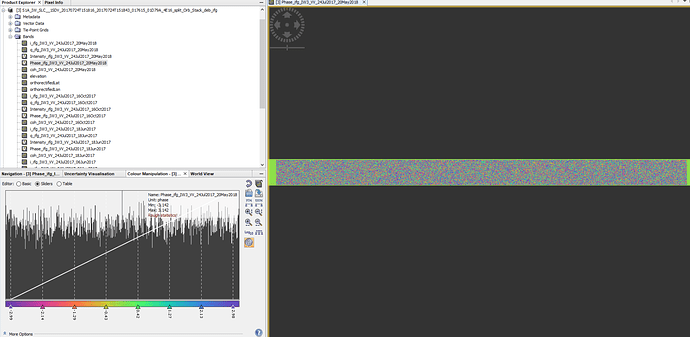
My suggestion is to check the output of the coregistration to see whether both master and slaves seem fine, as well as the interferograms.
I would like to ask you about the computer RAM resources and snap configuration employed to run the TOPSAR-coreg … as it could be the origin of all.
But it could happen that I am totally wrong…
Please check it out and let us know.
Hello Dear @mdelgado ,
Thanks for reply and contributing! I’m thinking the same thing with you that the problem is based on SNAP (maybe because of images, maybe some process in SNAP). My images i used are 5,8,9,6,4,7 (see Picture below). And i upload some screenshots about how some process looking as an image. Doing all process i m using 2 machine.
-
I’m working by intel Core i7 processor with 16 GB RAM supported SSD for SNAP analysis on Windows 10 system.
-
I’m working by intel core i5 processor with 4 GB RAM non supported SSD for other analysis(StaMPS-Matlab) on Linux 19.04 system. Best regards.
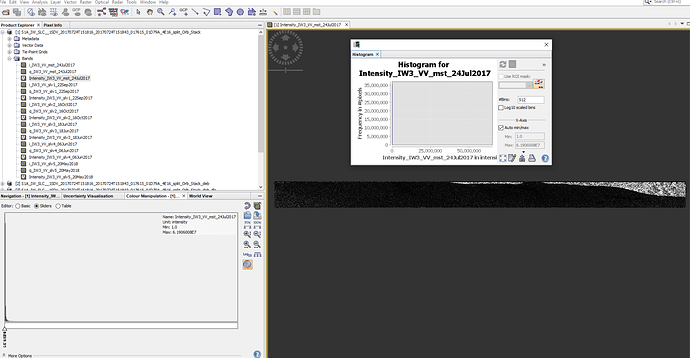
Well, amplitude images seems fine… but still strange the inf mean amplitude got from mt_prep_snap as said 0 pixels had non-zero values.
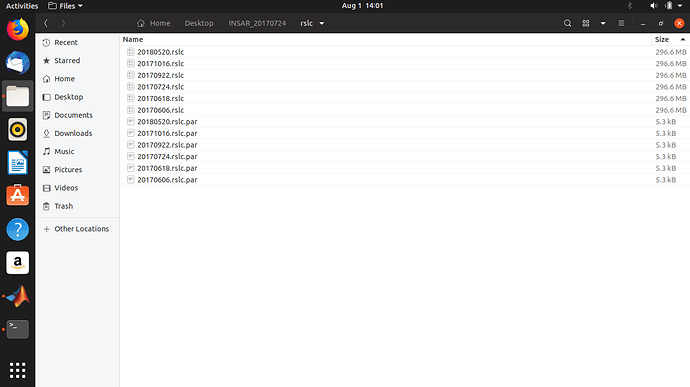
Maybe you got error while exporting for StaMPS PSI? Can you open the rslc files? It is better to do one more check than one check less.