Hi ,
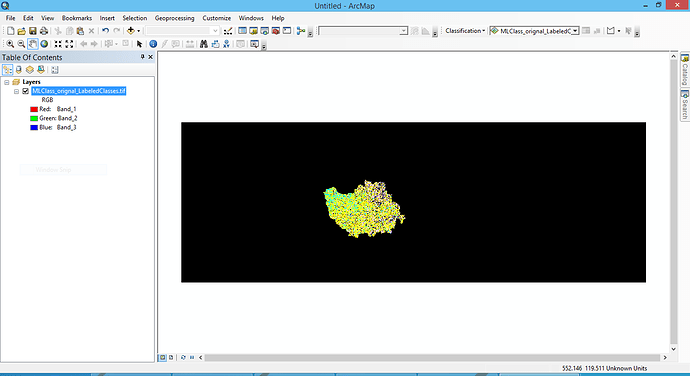
I have a labelled classified data of the Sentinal -1 image after the maximum likelihood classification. I saved it in tif format as well. However, when I try to load this Data in ARCGIS, i don’t see the labelled information.
I want to do the accuray prediction of the classifier.
Thanks
the labelling of the classes is something in the metadata of SNAP. This information is lost when you convert the data to GeoTiff. The raster only consists of integer numbers
It is not necessary to convert to GeoTiff: Please have a look at this related question: unsupervised classification
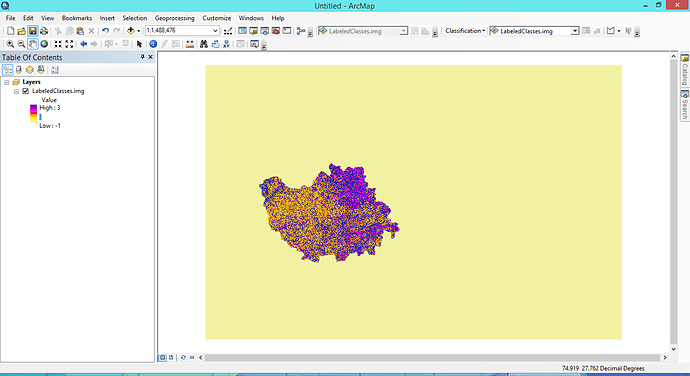
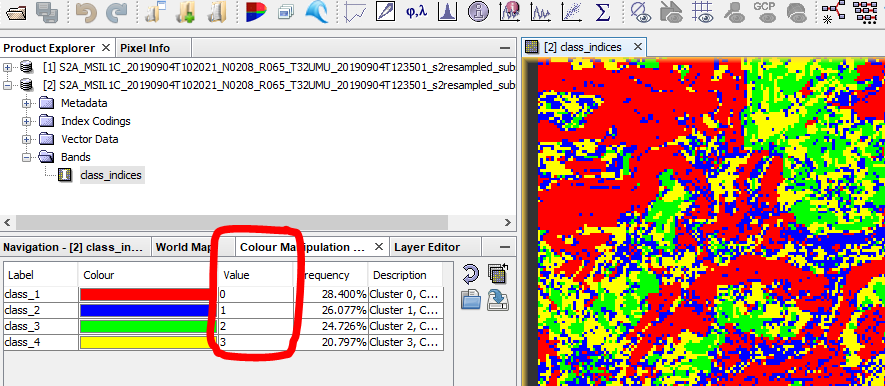
Thanks @ABraun for your reply. The image has been loaded with min as -1 and max as 3 value. I have 4 classes in my SNAP classifications ranging from 0-3. It is showing -1 as well .
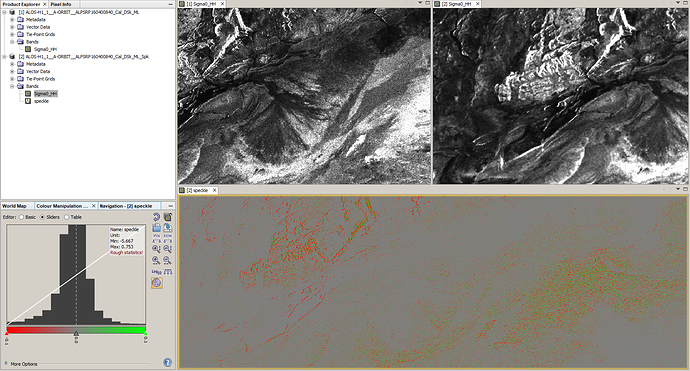
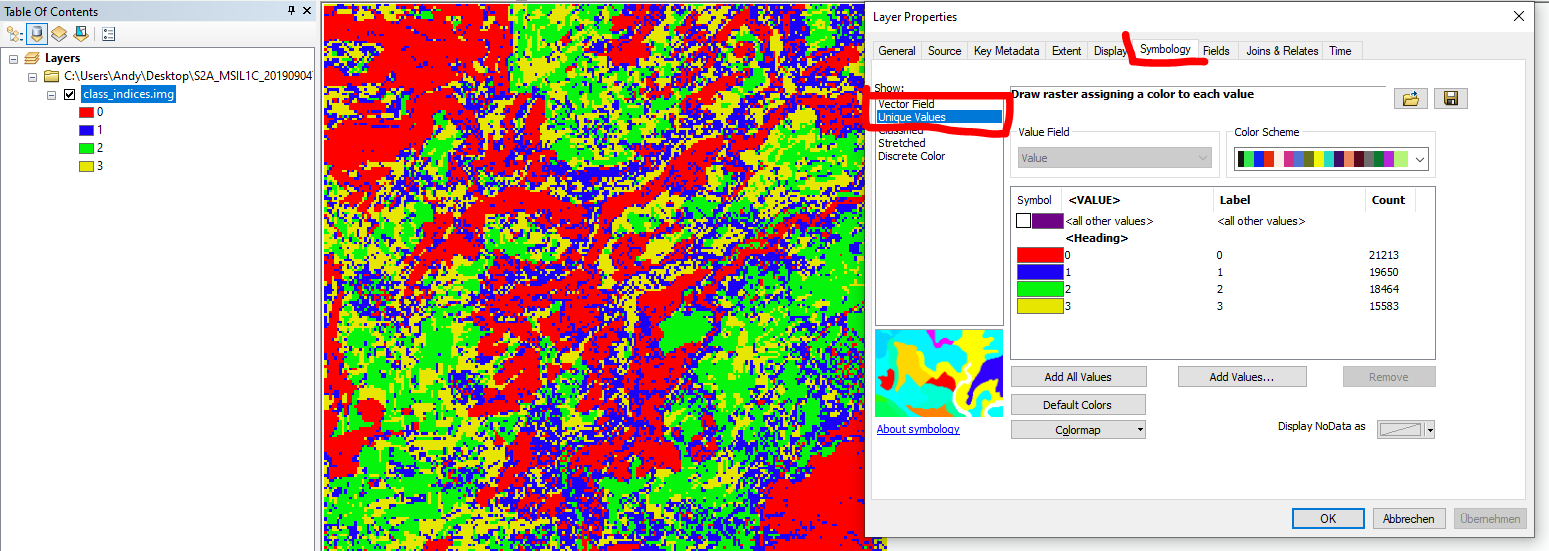
please check the actual range of the product in the image properties or make sure you have a unique color symbology instead of ramp:
Data in SNAP
Data in ArcMap
In your data -1 is probably the value outside the classified data.
yes it came after selecting this option. Thanks alot for your help.
I have taken the ROI using the sentinal 2 image and classified the images using maximum likehood algorithm in SNAP.
One image is the orignal sentinal 1 image terrain corrected.
The other image is the pre-processed sentinal 1 image with pre-processing( orbit file, calibration,speckle filtering lee filter,terrain correction)
Accuracy of orignal image= 67.2%
Accuracy of pre-processed image= 68.23%
There is not a huge difference in the results? I am wondering why is that happening?
Accuracy is almost same so why do we need to pre-process the image with so many steps.
Most of the preprocessing measures improve the quality of the image (geolocation, reduction of speckle) which are not directly visible in one single accuracy measure. The accuracy does not say much about speckle in the image, yet it makes sense to remove it before classifying. Similar for the calibration. Although you do not see the difference, it is good to reduce the impact of the incidence angle in your images so that similar pixels in near and far range are classified the same.
so how can we see how much amount of speckle is there before and after speckle filtering?can we implement it in SNAP.
there are several ways to check the efficiency of a filter
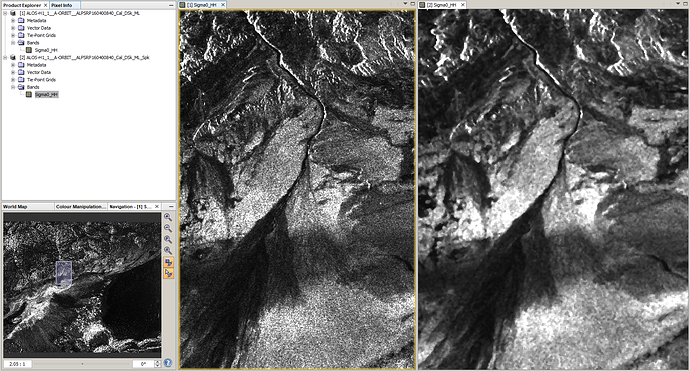
1. compare before and after using the window tiling options 
left: before filtering
right: after filtering
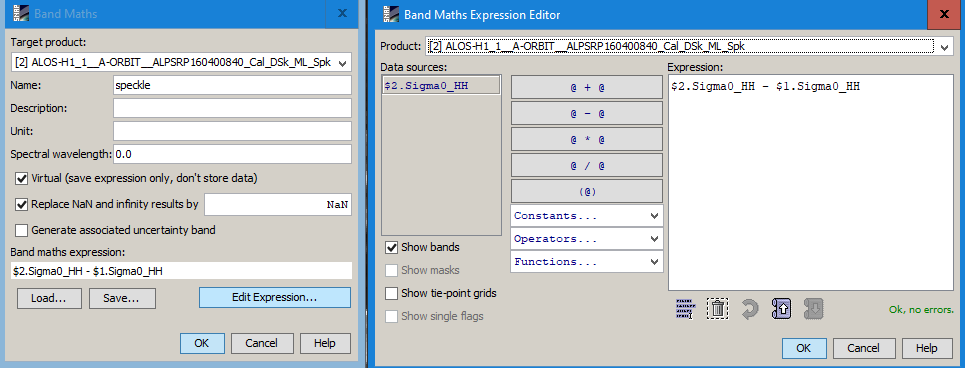
2 Calculate a difference image
Use the band maths to retrieve the amount of speckle which was suppressed during the filtering
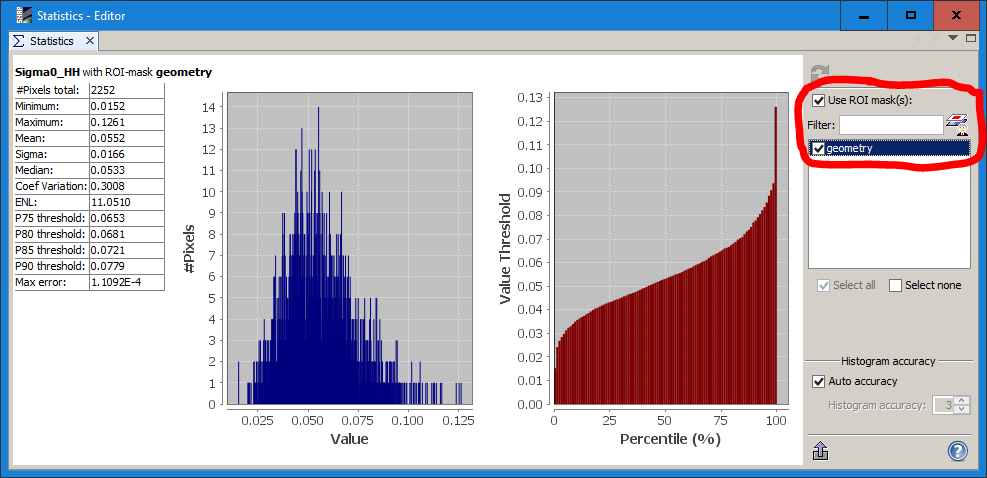
3 Determine the coefficient of variation
Digitize a AOI of a homogenous area
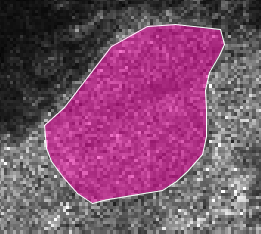

Apply the Speckle Filter
Now use the statistics tool to calculate the coefficient of variation for this area
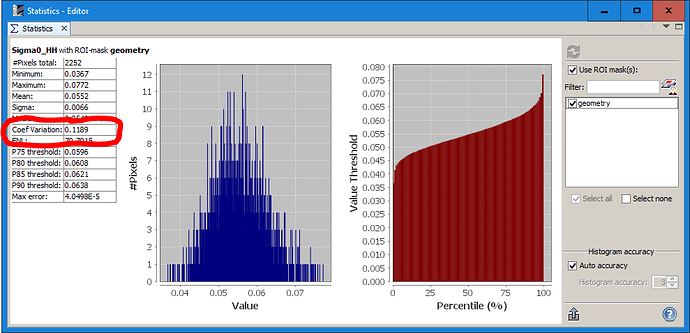
And the same for the product after filtering to see how much the variation decreased through the filtering
very helpful indeed in simple steps!! Thanks