Dear @ABraun
If we have two TanDEM-X pair (one in ‘18May in monostatic mode’ and another one in ‘29May in bistatic mode’), then it is possible to detect changes between them by using upper method that we discussed?
Thanks
There are two approaches:
- InSAR: Make one DEM per date using the TanDEM images and compare the absolute heights you derived.
- DInSAR: Coregister the HH band of one image per date (e.g. HH of master 18May and HH of master 28May), calculate their interferogram and perform topographic phase removal (using the ASTER mosaic) and calculate the difference with “phase to displacement”. This however only works for coherent processes. And I am not sure if the perpendicular baseline between the two dates is suitable.
Both approaches rely in correctly generated interferograms and high coherence in the areas that are of interest to you.
Yes. I did before with two bistatic pairs but I did not know it is also possible with one bistatic and one pursuitMonostatic. Thanks
Why don’t you just give it a try?
These studies used both modes: I haven’t checked if they just compare or if they use monostatic pursuit and bistatic combined .
https://ieeexplore.ieee.org/abstract/document/6049768
https://ieeexplore.ieee.org/abstract/document/6086975
You don’t seem very confident about your ideas? 
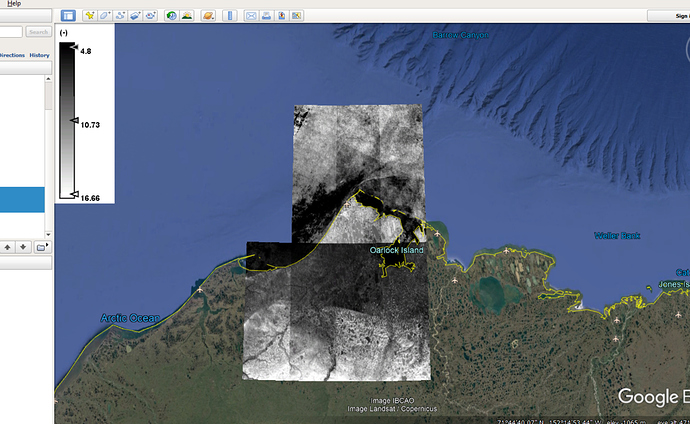
Based on our discussion, I did mosaicking to make an ASTER DEM over my area of interest but as you know, ASTER is a very flawed DEM and has lots of values over water (figure1). I want to ignore values over water.
I used mask option in SNAP but it did not work.
Any idea?
you could also try this one: https://www.eorc.jaxa.jp/ALOS/en/aw3d30/index.htm
But, similar to ASTER, it was based on optical data. If you mask out the sea areas before the terrain correction, you can select “mask out areas without elevation” as I mentioned above.
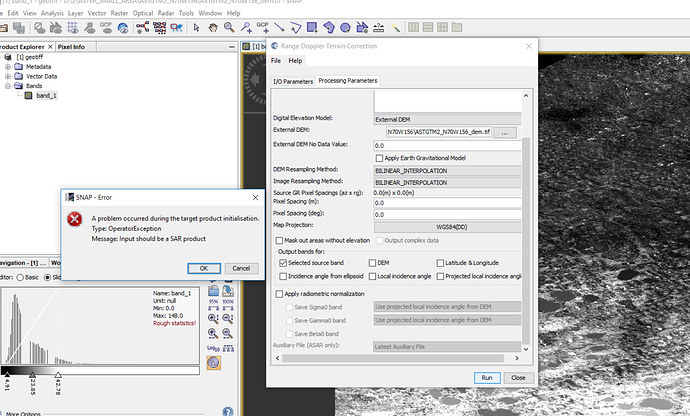
I think “mask out areas without elevation” is under ‘terrain correction’ tool and I tried it but it says it is possible only for SAR images but I want to apply on ASTER DEM.
Yes, applying terrain correction on the ASTER DEM doesn’t work. You have use the product after “phase to elevation” and just repeat the Terrain Correction step.
But as long as all pixels are valid in the ASTER mosaic, nothing will happen.
Dear @ABraun
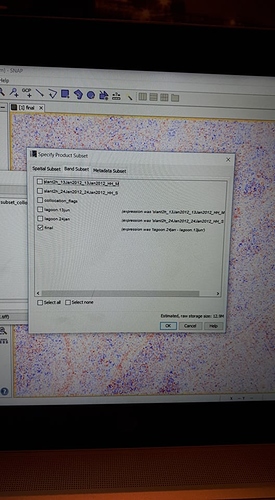
I again have same problem and although I converted bands and save them, again ask me below question.
Do you wish to include reference dataset into in your subset definition?
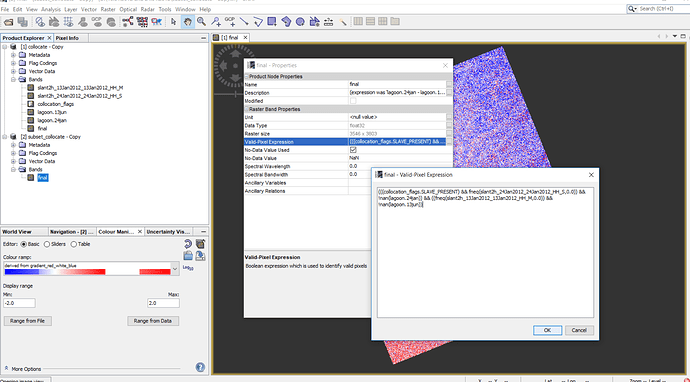
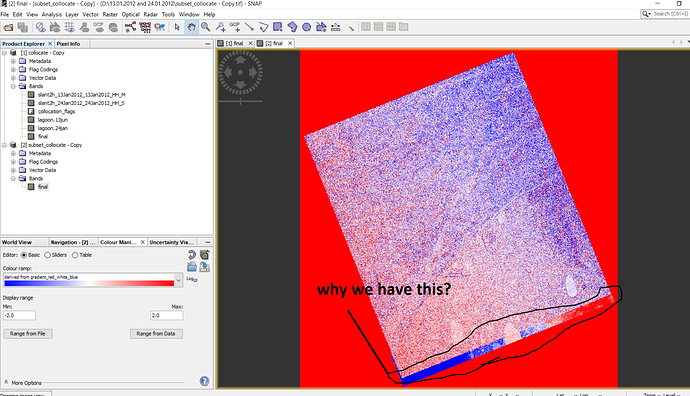
Then I chose ‘no’, result is saved but I cannot open the data but when I detete ‘valid pixels expression’ , then results will be opened but result has some extra part???
Figure1. difference map before extractiong in geotiff
Figure2. Image is not opened before deleting ‘valid pixel expression’
Figure3. We have some extra part after deleting ‘valid pixel expression’
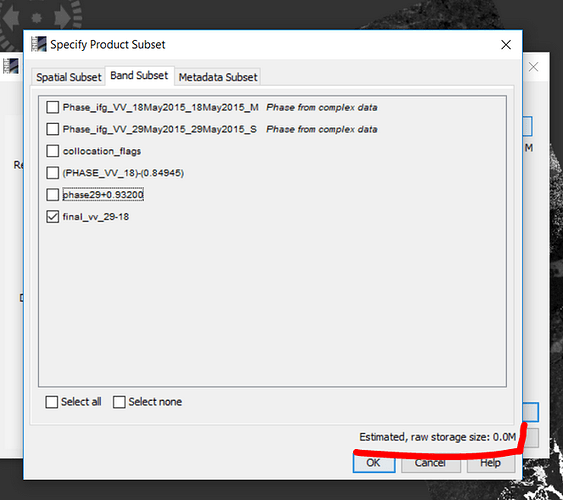
If the estimated raw storage size of your subset is 0.0 MB, your bands to export are only virtual.
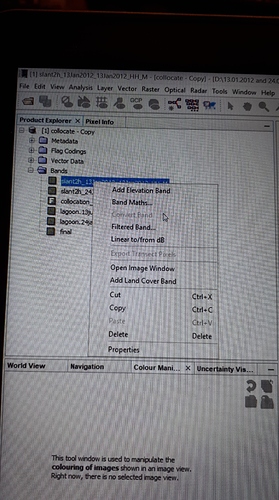
Right-click these rasters > convert band and save your product. Close the product and open it again. You can then export parts of the product (single rasters) without being dependent from a maths expression.
When I want to convert rasters, they look already converted.
Where can I find ‘the estimated raw storage size’?
Somehow, the products you want to export are dependent from the collocation flags. If they are not included, the output fails.
Is it an option to directyl work with the img files in the data folder instead of exporting GeoTIFFs? Both work fine in most GIS software.
Open the data in a GIS (QGIS, ArcMap,…) and save it as a GeoTIFF from there.
The product is saved in dim format and when I tried to open in QGIS, it was failed. I used ‘add vector layer’ and ‘add raster layer’ but they did not work.
use “open raster” and take the img file inside the data folder.
I did but when I open img in QGIS, then it has an extra part (look at attachment), that there is not in SNAP.
@ABraun…If you look at here, we had this problem after deleting ‘valid pixel expression’.
@ABraun
I had two DEM images for two different days.
- first DEM (3546*3803)
- Second DEM (3498*3713)
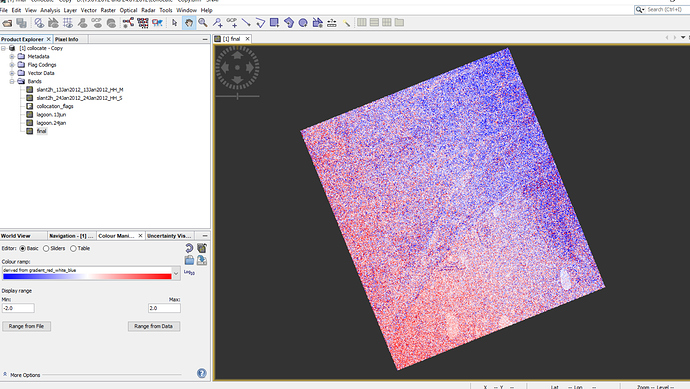
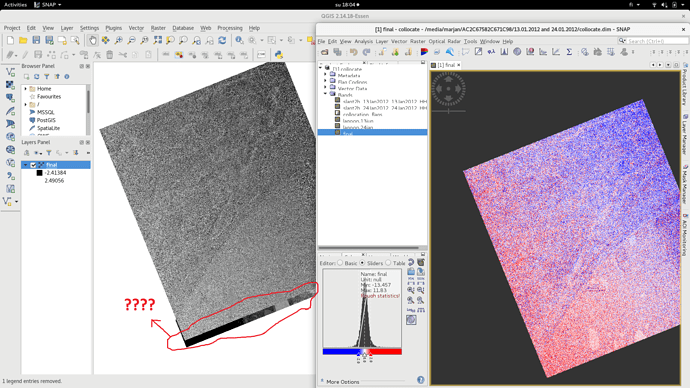
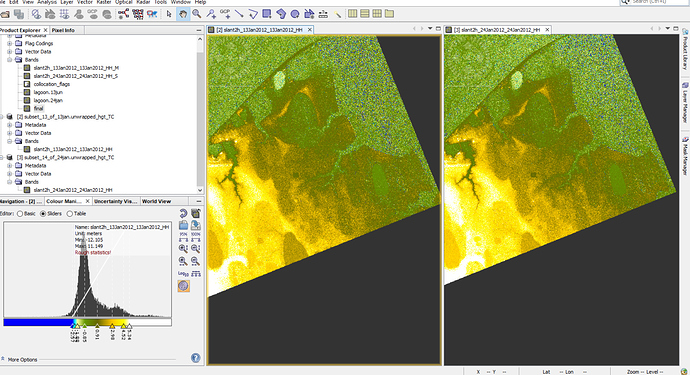
Although now pixels are same (3546*3803) after collocation for both images but one of them looks bigger that another one by eye (figure 1 ) but when I made difference map from them, everything was fine in SNAP (figure2) but when i deleted ‘‘valid pixel expression’.’ OR open difference map (img) in QGIS (figure3), then the extra part is obvious there.
figure1
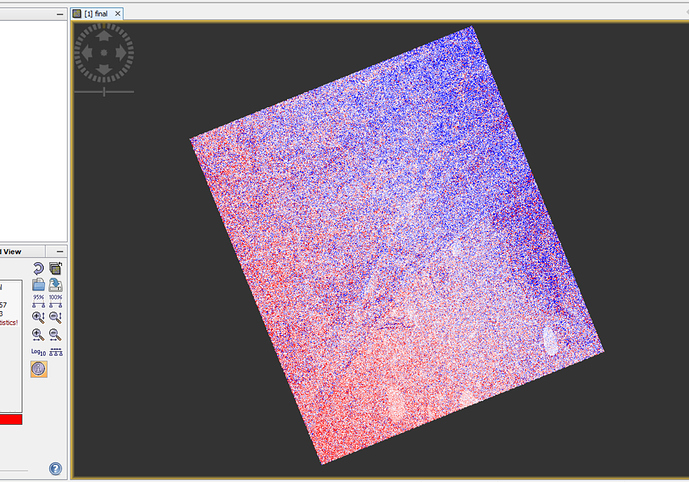
Figure2
figure3
you can crop this area by
- digitizing a polygon with the desired extent
(make sure it has the same spatial reference as the raster) and - using “clip raster by mask layer”
This writes a GeoTiff as you need it.
Here is a demonstration: https://www.youtube.com/watch?v=zAWA4NTw0Zs