Hello everyone,
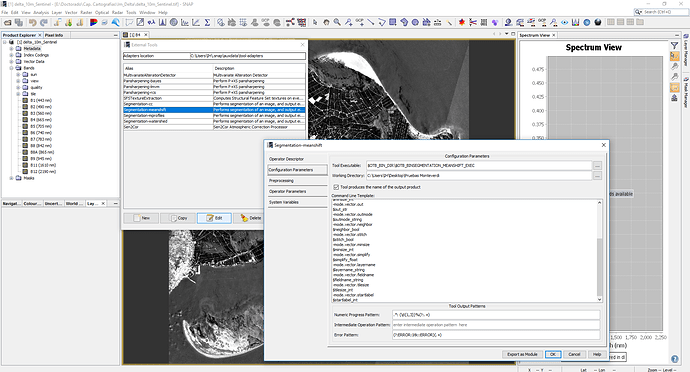
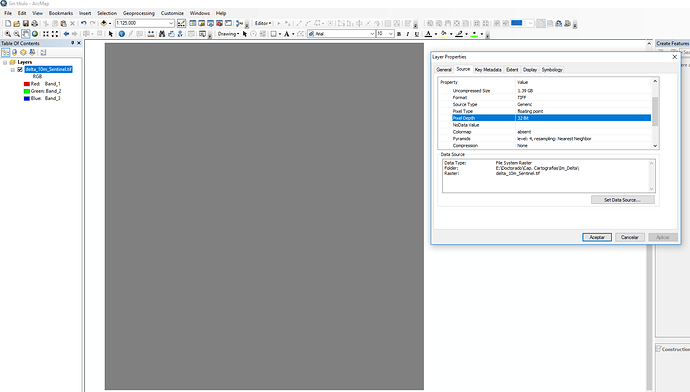
I’m having trouble with the Orfeo Toolbox segmentation tools from SNAP. Apparently the work fine, creating a shapefile in the indicated working directory. Nevertheless, that shapefile does not open. When I try to load the shapefile as a layer over the satellite image, it appears an error saying that couldn’t open file because a premature end of file. That same file does not open in ArcGIS.
I have followed all the threds I have found in this forum, but nothing seem to be quite the same as my situation, and none of the solutions seem to work.
When I do the segmentation directly on Monteverdi, it does work fine and I can open it as a layer in SNAP, so I think the Orfeo installation might not be the problem, and should be something related to the communication between SNAP and OTB.
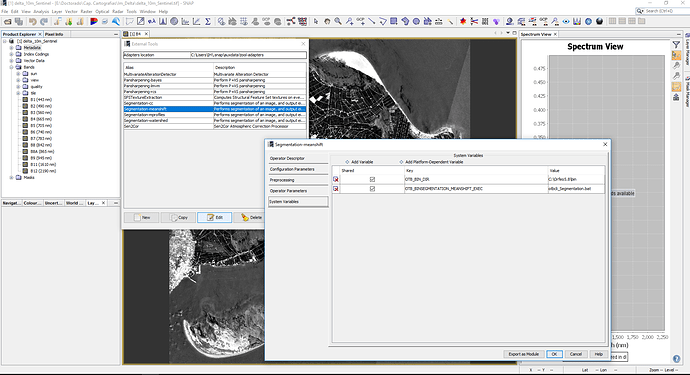
Something that caught my eye and maybe has something to do with it. When editing the external tools in SNAP, it never saves the configuration I have indicated in a previous sessions (I mean the working directory or the system variables) (it does save it during the session), and if I try to run the tool it gets me the same error shown here Segmentation on SNAP
I have reinstalled several times the external tool, the SNAP, the OTB, but nothing seem to work in that final step of finishing the .shp. The SNAP does not report any error, but does the whole process in about 20 seconds, which seems odd.
I also created the java folder in the SNAP folder as stated in here External Tools disappear when restarting SNAP - workaround
Do anyone have an idea of what could be missing?
Sorry for the long post, hope you can help me.
Thanks a lot,
Felipe
I attach some information about versions and logs in case they are useful:
SNAP 5.0
Orfeo 5.8.0 (I also tried with Orfeo 6.0 with same results)
Win 10 Pro
I’ve worked with two images, both GeoTiff:
One from Sentinel 2 downloaded from the SciHub portal. It was corrected with sen2cor, subseted to the area of interest and resampled to 10m spatial resolution with SNAP. Geographic system WGS84 UTM Zone 31N. Size 1.6GB.
The other from Landsat 8, downloaded directly from Earth explorer, the product of surface reflectance. Subset to the area of itnerest with SNAP. Geographic WGS84 UTM Zone 31N. Size 30 MB.
The log from the most recent session is here messages.log (125.7 KB)