yes. But as I said, it is important what the script prints.
I created a new directory and executed “mt_prep_snap” there itself.
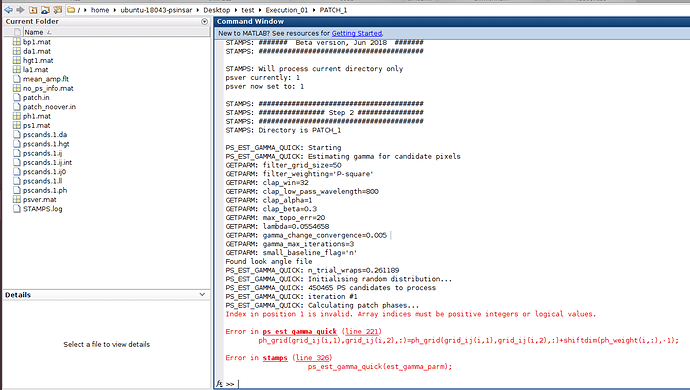
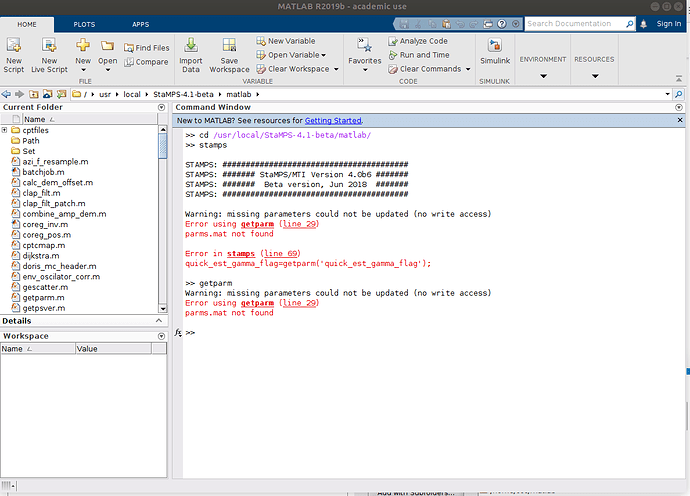
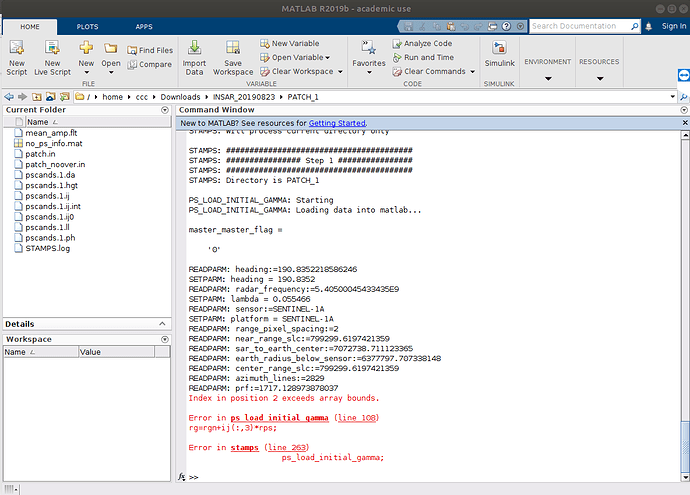
stamps(1,1) executed success fully,
stamps(2,2) getting an error as follows
how many interferograms are you using?
Have you visually checked if the interferograms produced in SNAP look alright?
Please also consider these discussions on the same (or similar error):
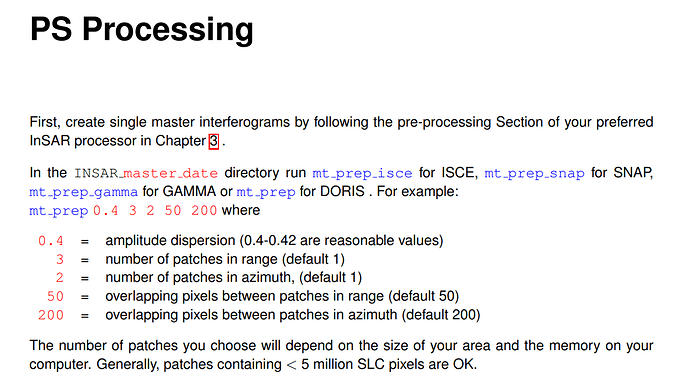
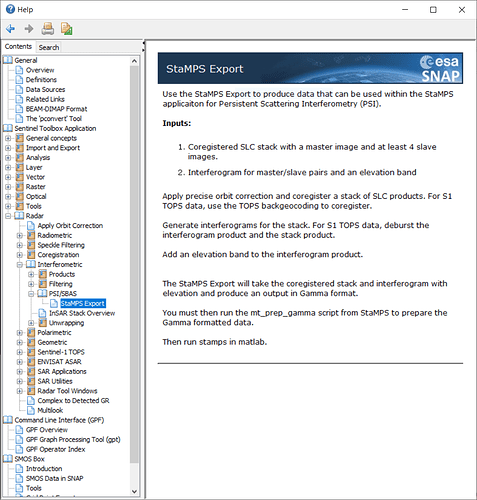
in the SNAP help, it is showing to execute “mt_prep_gamma” script
in the manual, it is showing that, execute “mt_prep_snap” script.
which is the correct one to be followed?
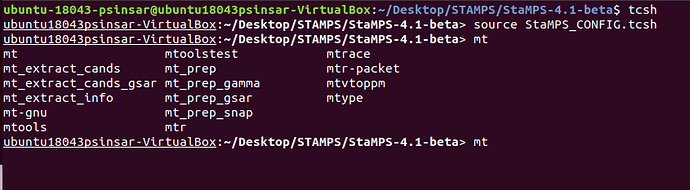
“mt_prep_gamma” is showing in tcsh enabled terminal
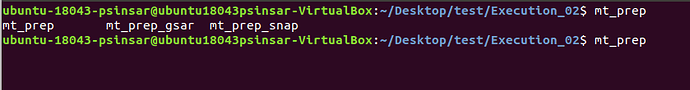
in the terminal from execution folder, “mt_prep_gamma” is not showing.
I am using 22 Interferograms.
All are looking fine while opened in SNAP.
mt_prep_gamma was the one initially to be used, but it did not work for Sentinel-1 data very well.
Therefore, they released a second one (mt_prep_snap) at a later time which should be used now.
I recommend, creating a new work folder and running this script in there again. The files exported from SNAP can be maintained.
Hi,
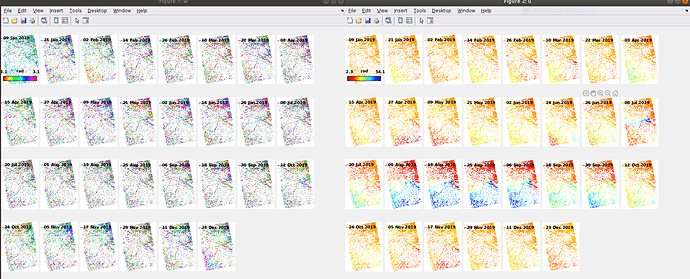
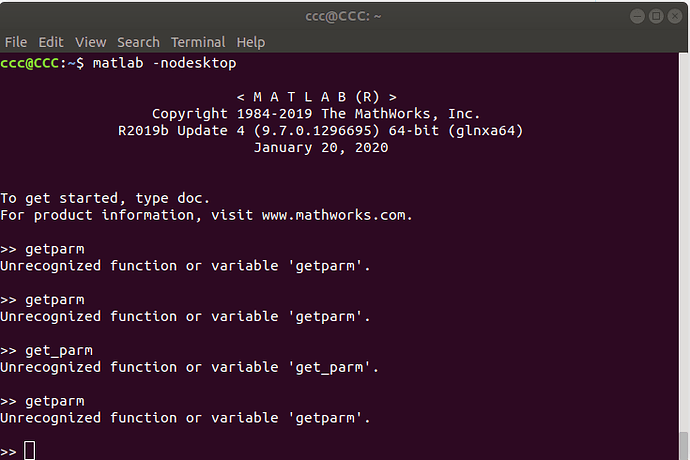
I have processed 30 scenes and obtained as follows.
How to analyse the results scientifically …!
w and u are not the most important plotting options. If you want to measure surface deformation, you need v, at best in combination with d and o.
The ts option also allows you to plot the displacement of a selected area.
It depends on your research question how you want to to analyze the results.
Extract from the manual
'w' for wrapped phase
'w-d' for wrapped phase minus smoothed dem error
'w-o' for wrapped phase minus orbital ramps ('w-dm', 'w-do', 'w-dmo')
'p' for spatially filtered wrapped phase
'u' for unwrapped phase
'u-d' for unwrapped phase minus dem error
'u-m' for unwrapped phase minus and master AOE
'u-o' for unwrapped phase minus orbital ramps
'u-a' for unwrapped phase minus topo-correlated atmosphere
('u-dm', 'u-do', 'u-da', 'u-dmo', 'u-dma', 'u-dms', 'u-dmao', 'u-dmos')
'usb' for unwrapped phase of small baseline ifgs
('usb-d', 'usb-o', 'usb-a' also 'usb-do','usb-da', 'usb-dao')
'rsb' residual between unwrapped phase of sb ifgs and inverted
'd' for spatially correlated DEM error (rad/m)
'm' for AOE phase due to master
'o' for orbital ramps
's' for atmosphere and orbit error (AOE) phase due to slave
'v' mean LOS velocity (MLV) in mm/yr ('v-d', 'v-o', 'v-a', 'v-do,
'v-da', 'v-dao')
'vs' standard deviation of MLV (mm/yr) ('vs-d', 'vs-o', also 'vs-do')
'vdrop' MLV calculated from all but current ifg (mm/yr)
'vdrop-d'
'vdrop-o' (also 'vdrop-do')
These might also help:
sorry, could you explain this step more details, please. Because I got the same problem also.
Thank you
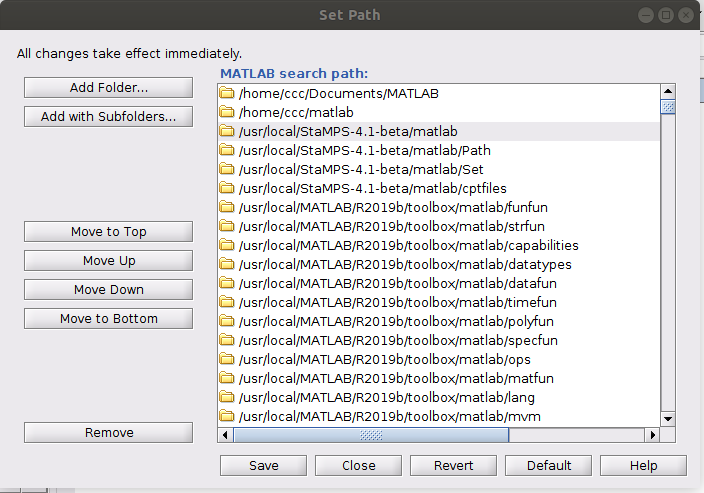
MATLAB needs to know the folder where the stamps scripts are located. You can do this in the GUI (by adding this directory to the list of folders by clicking on “Set Path”.
Another option is to add this folder to the line where the MATLABPATH is defined in the StaMPS_CONFIG file
(line 45 here: https://github.com/dbekaert/StaMPS/blob/master/StaMPS_CONFIG.bash)
seems that your data is located in a directory with restricted writing permissions.
I already change the permission, but no change with the error
please navigate to the directory where you store your files processed by mt_prep_snap
You have to execute the matlab commands/scripts there. If they are included in the Path list, they are found from any location.
I guess the StaMPS_CONFIG.sh is not in the src folder, but in the parent one.
Can you check?
according to the last screenshot he tried both
I just assume the name he tried is not correct
Please try StaMPS_CONFIG.bash 
According to the github stamps file list, it should be call like this, and found in the parent folder, not inside the src
Oh right. 
Had you used the updated ps_load_initial_gamma.m file I provided in the forum (somewhere around March 2019)?
I would appreciate you search for it in the forum, from my cellphone I can’t find it.