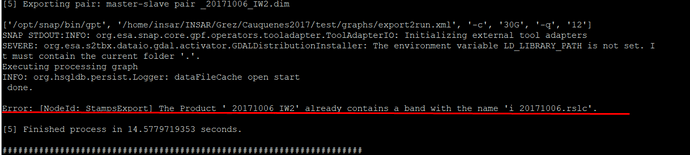
Hello, I have seen that other people have this same error, but have not found the solution. This occurs by running the stamp_export.py script and only with the master image.
I don’t know if you have any ideas that you can help me with.
I look forward to your responses.
Regards.
Did you prepare the master manually?
If so, please make sure that you have only selected VV polarization in the TOPS Split operator.
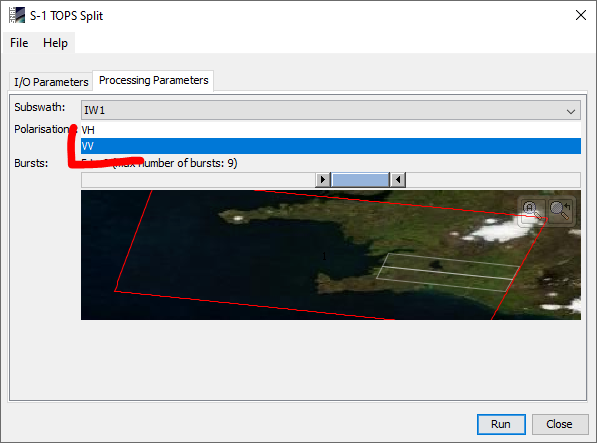
The final stack should only consist of VV bands (of master and all slaves)
When using snap2stamps it does this automatically by assigning the master image in project.conf, right?
the master image is referenced in the config file, yes. But you still have to prepare the master, manually as recommended here, to make sure it is processed correct.
Also, the master should be stored in a separate folder so that the coregistration will not use it as a slave. This would lead to a stack with identical master and slave date (probably also one cause of the error)
I will follow your recommendations, thank you very much and sorry for the inconvenience.
no need to apologize. In case one of the suggestions fixes your error, please report in here so others can learn from this case.
Indeed I had the master image with the slaves, now everything works without error.
That’s how I solved it:
from the stack_deb just delete the duplicates of i_mst,q_mst and Intensity_mst, and the error will no longer appear and you can export the pair made with Master-Master
Obviously this step is if you do it manually and not using xml or script_python 
hi, I’m adding this topic conversation because I am having a similar problem with coregistration of two ALOS-2 slc images and this is the only place I saw it mentioned. I have used the coregistering procedure many times with different ALOS-2 images, however, about a week ago the msg in the graphic started appearing when executing the read tab in the coregistration procedure. The band mentioned in the error is an earlier product of processing the two images (a month or more ago) but that product does not reside on the workstation at this point. the folder names differ from those used previously. I put the master and slave in separate folders, however, the error persists. I convert the bands saving each step before processing. I have also reloaded the original slc images in case the original files were corrupted, however, the same msg occurs. the final product is the interferogram. thanks
are these dual-pol data?
hi, yes, HH, HV. I did not subset the bands. thanks
please first use “Select Band” to reduce your images to HH before you coregister them. This not only saves time and disk space, but also prevents these naming errors. The cross polarization channel does not bring interferometric benefit for most InSAR approaches. Effect of polarization on interferometric products
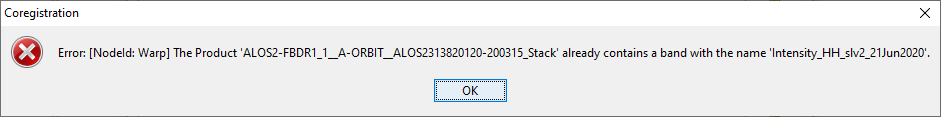
hi ABraun, I subset the two images and included the Band Select to just keep the HH band. I got the error shown on coregistration screen shot below… 
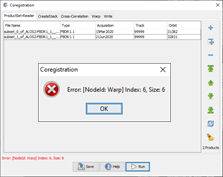
can you please include larger screenshots?
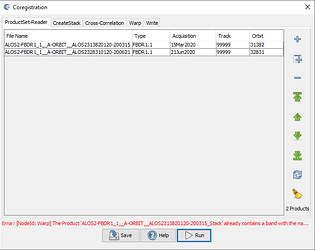
sorry, the larger screen shots attached…
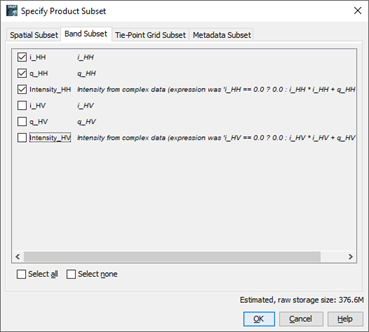
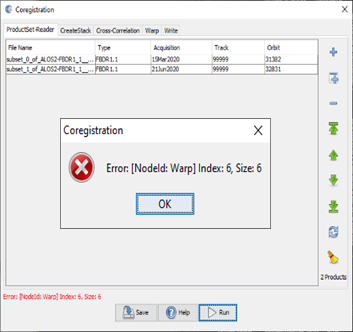
have you stored both images to BEAM DIMAP forma after creating the subsett?
hi ABraun, yes, both were subsets were saved as dim files
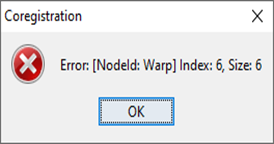
And which parameters are selected under Warp?
hi, ABraun, the coregistration failed at the product read tab. I’m not sure why the error includes “Warp.”
Actually, the message states that the error comes from the Warp tab, but it is displayed already when you open the data so it could be related to the metadata. Please show the Warp tab