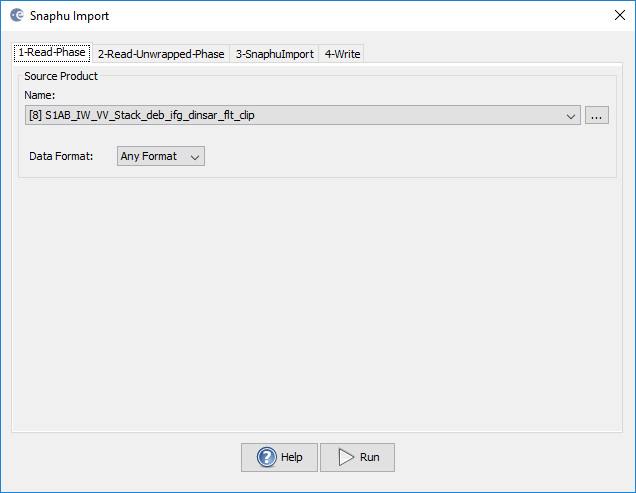
did you select the right reference file (1-Read-Phase) in the import module? This should actually copy the source image’s metadata into the imported unwrapped phase.
1-Read-Phase (this is the reference file, usually the one before the unwrapping)
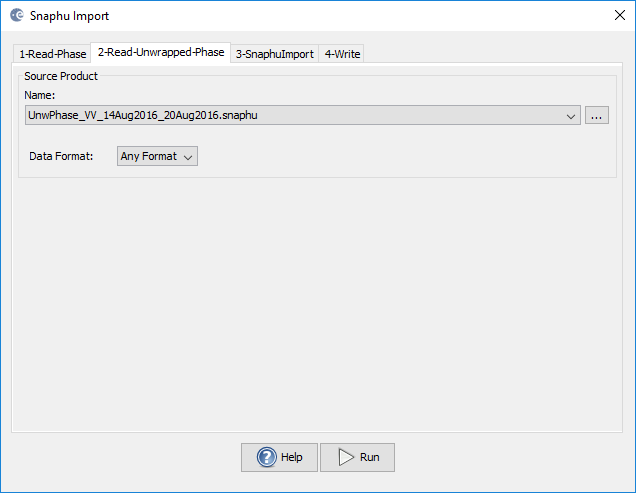
2-Read-Unwrapped-Phase (here you chose the hdr file of the unwrapped phase created by snaphu)