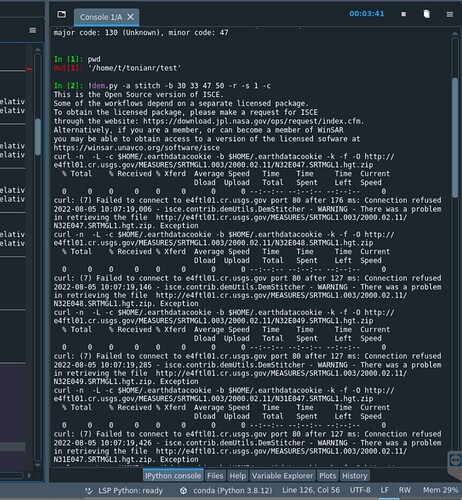
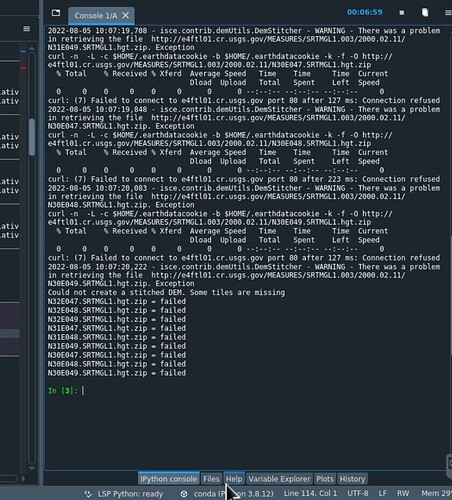
I got this error before for something simple. It could be something very simple like what variable you list first in the input file. Reorder the listing so that azimuth looks is listed before range.
Looks like you have sentinel data, heres one of my sentinel input files:
source_data slc_stack
slc_stack_path /home/toni/WCF_SENTINEL_processing/S1A_RO_121/merged/SLC/
slc_stack_reference 20180724
slc_stack_geom_path /home/toni/WCF_SENTINEL_processing/S1A_RO_121/merged/geom_reference/
slc_stack_baseline_path /home/toni/WCF_SENTINEL_processing/S1A_RO_121/merged/baselines/
maskfile /home/toni/WCF_SENTINEL_processing/S1A_RO_121/merged/geom_reference/shadowMask.rdr
azimuth_looks 1
range_looks 3
aspect_ratio 3
lambda 0.056
slc_suffix .full
geom_suffix .full
Dear Tonianr, I appreciate your help.
I got your error too but I didn’t get how you solved it, can you tell me how can i solve it?
my error is in stamps(1,1):
Subscripted assignment dimension mismatch.
Error in ps_load_initial_isce (line 144)
ph(:,i)=complex(ph_bit(1:2:end),ph_bit(2:2:end));
Error in stamps (line 285)
ps_load_initial_isce(data_inc)
I’m not sure what is causing that error but you can check that your patch directories do not have empty files (0 bytes). You can also check the https://groups.google.com/g/mainsar forum for the error.
Oh yes, exactly it is 0 bytes. So I should decrease the number of mu patches?
hi, dear Tonianr,
I am working in ISCE software,
I want to download Dem but I can’t download.
I have an error:
isce.contrib.demUtils.DemStitcher - WARNING - There was a problem in retrieving the file http://e4ftl01.cr.usgs.gov/MEASURES/SRTMGL1.003/2000.02.11/N30E048.SRTMGL1.hgt.zip. Exception
would like help me please.
Dear Tonianr, Could you give me a sample of prep_isce.py for TSX data please?
and is there any instruction for using TSX in isce?
Hi, I had that same problem 1 week ago, seemed to have been a problem with the website. It works now!
Hi Nadiaha,
There are no instructions for using TSX in ISCE StripMap stacking. Is there a particular step you need help with?
Here is a sample of my unpack_TSX.py and prep_TSX.py scripts (from my anaconda3/envs/isce2/share/isce2/stripmapStack directory).
I use this command in python to unpack my Starring Spotlight TerraSARX (TSX) images:
!prep_TSX.py -w ../ -i downloads/ -f /T*X1_ST_021*
before using prep_TSX.py make sure to unzip your files, I unzipped my tar.gz files using:
os.chdir('/downloads')
!cat ../targz/*.tar.gz | tar -xvzf - -i #unzip
this unzips them to the downloads directory.
prep_TSX.py (1.8 KB)
unpackFrame_TSX.py (1.6 KB)
Hope this helps!
hi dear,
I can’t download Dem yet
could you help me,please
Dear Tonianr, How could I make a stripmapStack directory in my isce2 environment?
also I found you use TX1_ST_021, is this name like “TSX1_SAR__SSC______SM_S_SRA_20190420T024646_20190420T024654”?
Hi dear
this is my problem . I’m using a ISCE2.
my study area is 30,33:47:50
You will need to find the stripmapstack directory in your isce2 documents
dem.py -a stitch -b 28 29 -83 -82 -r -s 1 -c #SNWE
Is th command I use in python
I use exactly the same command, but I still can’t download. Can you test this region for me,please.
dem.py -a stitch -b 30 33 47 50 -r -s 1 -c
Dear Tonianr, thanks for your help I found my stripmapstack dir, and used this command for unpack my data but i got error:
unpackFrame_TSX.py -i home/Desktop/Nadiaha/data/ -o home/Desktop/Nadiaha/data/TSX1.SAR.L1B/TSX1_SAR__SSC______SM_S_SRA_20190420T024646_20190420T024654/TSX1_SAR__SSC______SM_S_SRA_20190420T024646_20190420T024654.xml
Could you tell me what wrong is in my command?
I suggest using the scripts as described you need your unzipped TSX files in a directory called “downloads” and an available “SLC” directory. Also I’m not sure if that’s an error message or is that the command you used? Prepare Stack to Stamps with TerrasarX data (ISCE) - #35 by tonianr
Looks like something is going wrong with the server. You’d need to take this issue to the ISCE2 discussions forum on GitHub, to see if they’re any solutions!
Dear Tonianr, i unzipped all my tsx data in a folder which its name is downloads and i run the below command in downloads directory and i got the following error:
!prep_TSX.py -w …/ -i download/ -f /TSX*
Traceback (most recent call last):
File “/usr/local/ISCE/isce2-main/contrib/stack/stripmapStack/prep_TSX.py”, line 29, in
if not os.path.exists(file_list[0]): #checking for atleast one file
IndexError: list index out of range
Could you help me to fix it?