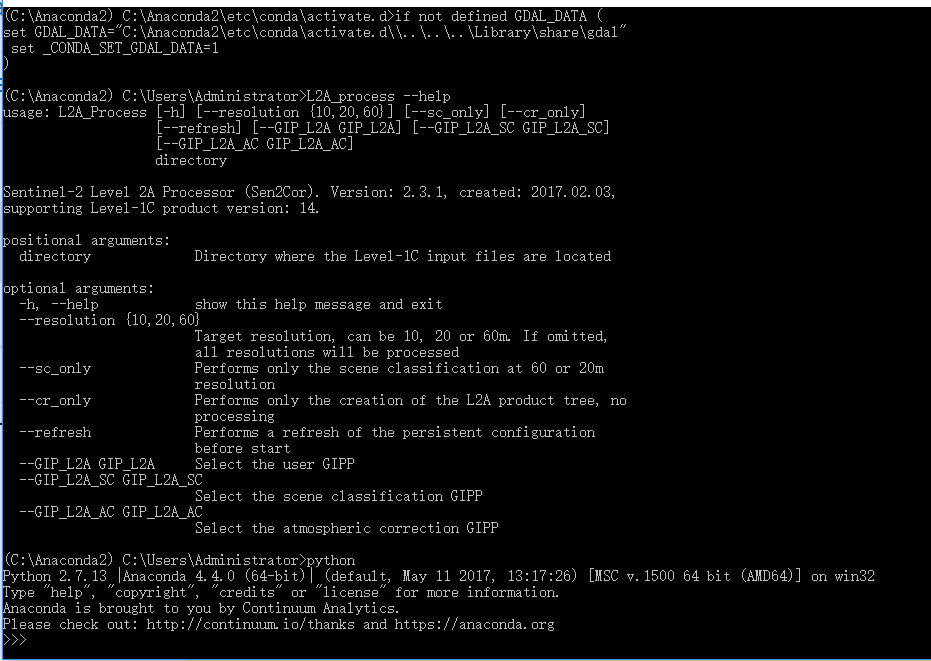
and when i type Python in cmd, i get this
that’s good on the one hand because sen2cor uses the correct python version,
but bad on the other hand becuase it doesn’t find gdal.
A solution was proposed here:
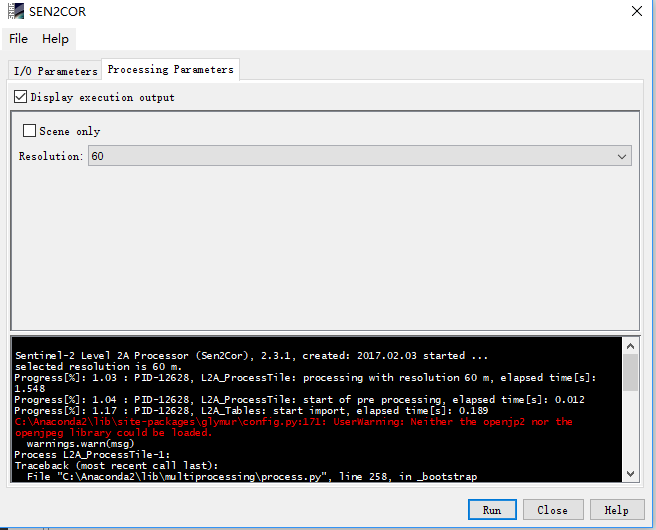
hello,I want to ask a question.when I use the sen2cor. It shows:
I want to ask what I should do? where is wrong? thank you!
please have a look at these topics:
thank you for your reply. these question is about 64 bit python, but The python in my computer is 64 bit. I also found this problem.So I just want to ask what I should do? thank you!
according to your screenshot, everything works fine.
Saludos para todos.
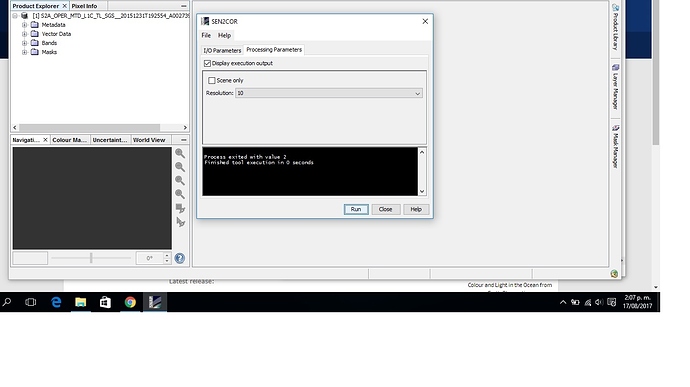
Me aparece lo siguiente al ejecutar sen2cor
Process exited with value 2
Finished tool execution in 0 seconds.
.
I would appreciate any clarification or assistance.
Saludos para todos.
please enter “Finished tool execution in 0 seconds.” in the search field and you will find lots of topics with helpful posts.
Greetings, thanks for replying.
I use windows 10 64-bit, I was consulting the resources in the link that you left me and I think I have solved that problem, change the value of SEN2COR_HOME, update the plugimn of sen2cor from SNAP and download from anaconda “glymur” because it present a problem with JPEG2000.
But now it appears to me
L1C user product directory must match the following mask: S2A_MSIL1C * .SAFE
But is: S2A_OPER_MSI_L1C_TL_SGS__20151231T192554_A002739_T18NWL_N02.01
Process exited with value 1
Finished tool execution in 5 seconds.
Sorry for the inconvenience.
Hi @Gabriel,
it seems that you are trying to run sen2cor using as input a granule, you have to use as input a L1C full product.
Hi,
After a quick look I saw that nobody answered your question… I also dont have an answer but I am facing the same issue…
After batch processing around 100 images with Sen2Cor I noticed that the cache folder took over my disc space… I was also wondering how this can be avoided and if the cache could be deleted.
Cheers
Lazaros
Hai Ralf,
I am trying to install sen2cor from two weeks but not able to do that. Can you please help me to install sen2cor and make it working on my PC using teamviewe software. Please help me out.
Hello!
I´m using Sen2cor in SNAP and I´ve the same problem in my two computers…
In the middel of the process…
File “C:\Anaconda2\Lib\site-packages\sen2cor-2.4.0-py2.7.egg\sen2cor\L2A_ProcessTile.py”, line 205, in process
if(sc.process() == False):
File “C:\Anaconda2\Lib\site-packages\sen2cor-2.4.0-py2.7.egg\sen2cor\L2A_SceneClass.py”, line 830, in process
if(self.tables.sceneCouldHaveSnow() == True):
File “C:\Anaconda2\Lib\site-packages\sen2cor-2.4.0-py2.7.egg\sen2cor\L2A_Tables.py”, line 2461, in sceneCouldHaveSnow
from PIL import Image
File “C:\Anaconda2\lib\site-packages\PIL\Image.py”, line 56, in
from . import _imaging as core
ImportError: DLL load failed: No se puede encontrar el módulo especificado.
I think I´ve installed everthing without any error…
any idea?
thank you very much!
Oscar
well,
I used the Standalone version and I could finish the process…
I hope we can use the snap6 and sen2cor soon without problems!
regards!
You need to mention the path of python.exe. With me it is Anaconda Installation. C:\Anaconda2\python.exe
Dear All
I am having some problems opening corrected output image from Sen2Cor in SNAP. I get the following error:
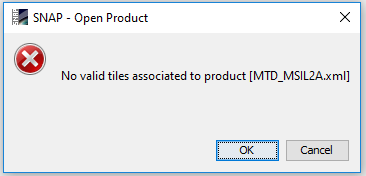
The image was corrected using Sen2Cor 2.5.5 and I am using Snap 6. I am also concerned about the errors in Sen2cor, Windows 10 (using PowerShell)
Should I be worried about these errors?
Hallo MahlatseK,
you should be worried about these errors since you cannot open and analyze your result. I assume you tried processing at 10 m spatial resolution only. If, then you should try to process again for all spatial resolutions. You may have a look to the problem report Sen2Cor-02.05.05-win64 - AttributeError: 'L2A_Tables' Object has no attribute '_L2A_Tile_PVI_File' where this is discussed in more detail. I hope this will help you and solve your problem as a workaround until the bug is fixed.
Thank you very much for the advice, I can confirm that I managed to run sen2cor without no errors following steps provided. However I am still unable to open the product in SNAP 6 getting same error as above.
Hi @MahlatseK,
have you updated SNAP? The new format is supported since version 6.0.1.
I have updated now its working like a charm  Thank you a lot.
Thank you a lot.