@JeroBande is right, the problem is the compiler version. StaMPS has to be compiled with gcc-7 (gcc-9 was installed in my case and used for the compilation of StaMPS which led to incorrectly compiled scripts)
So this is what it solved the problem in my case (partially copied from here). I cannot grant that it will solve all problems, but changing the compiler makes a difference. If you get stuck at one of these steps, please try to continue searching the web, because I have no clue how to troubleshoot most of these steps in case something goes wrong.
Adding ubuntu-toolchain-r/test PPA
sudo apt install software-properties-common
sudo add-apt-repository ppa:ubuntu-toolchain-r/test
Install several compilers
(we add 7, 8 and 9 so we have a list to chose from)
sudo apt install gcc-7 g++-7 gcc-8 g++-8 gcc-9 g++-9
Configure alternative compilers
sudo update-alternatives --install /usr/bin/gcc gcc /usr/bin/gcc-9 90 --slave /usr/bin/g++ g++ /usr/bin/g++-9 --slave /usr/bin/gcov gcov /usr/bin/gcov-9
sudo update-alternatives --install /usr/bin/gcc gcc /usr/bin/gcc-8 80 --slave /usr/bin/g++ g++ /usr/bin/g++-8 --slave /usr/bin/gcov gcov /usr/bin/gcov-8
sudo update-alternatives --install /usr/bin/gcc gcc /usr/bin/gcc-7 70 --slave /usr/bin/g++ g++ /usr/bin/g++-7 --slave /usr/bin/gcov gcov /usr/bin/gcov-7
Select preferred compiler
(here we want gcc-7)
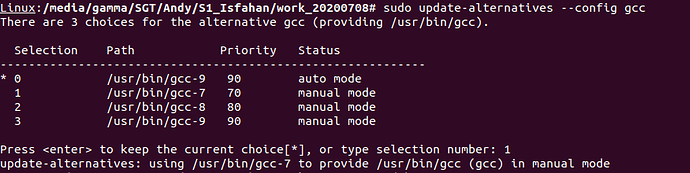
sudo update-alternatives --config gcc
this prints this table
There are 3 choices for the alternative gcc (providing /usr/bin/gcc).
Selection Path Priority Status
------------------------------------------------------------
* 0 /usr/bin/gcc-9 90 auto mode
1 /usr/bin/gcc-7 70 manual mode
2 /usr/bin/gcc-8 80 manual mode
3 /usr/bin/gcc-9 90 manual mode
Press <enter> to keep the current choice[*], or type selection number:
This means, that gcc-9 is used to compile code unless another is selected. To confirm that we want to use gcc-7 for the compilation of StaMPS, type
1
and hit Enter
Now, without closing the command shell, nagivate to the StaMPS directory and inside the src folder
type
make
and then
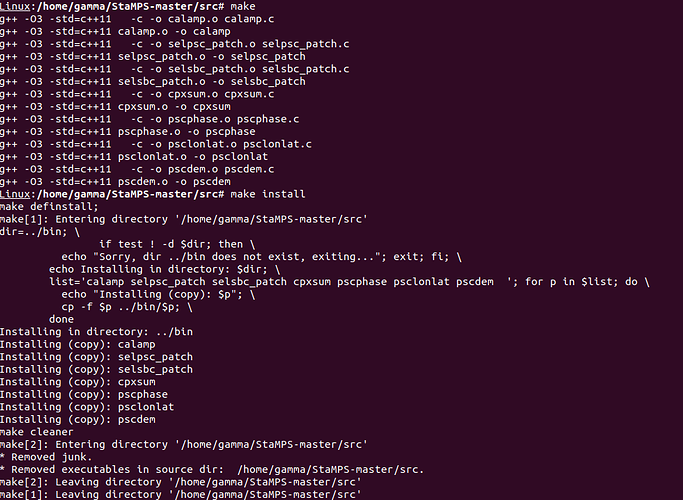
make install
This will compile all scripts required for PS processing. It should look like this
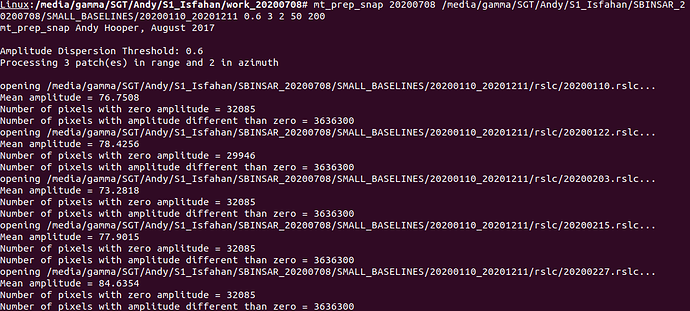
Now, the Segmentation fault (core dumped) error will no longer occur when you run mt_prep_snap again. I recommend to run this command in an empty folder (not the same as the INSAR folder exported by SNAP and also using the tcsh instead of the standard command line. Also, don’t forget to modify and source StaMPS_config.tcsh before you can run all scripts.
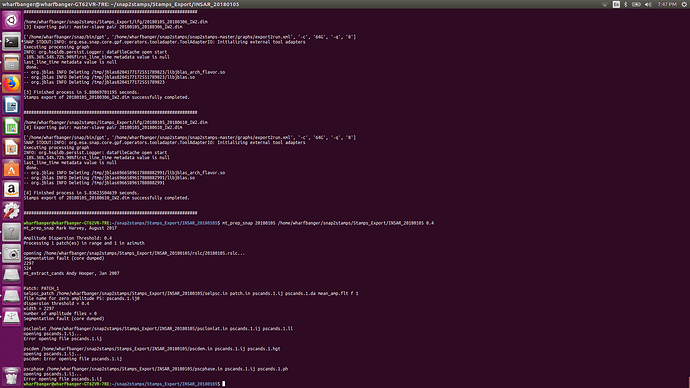


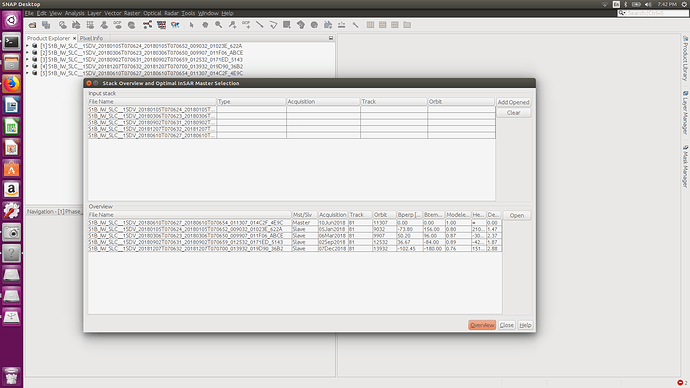
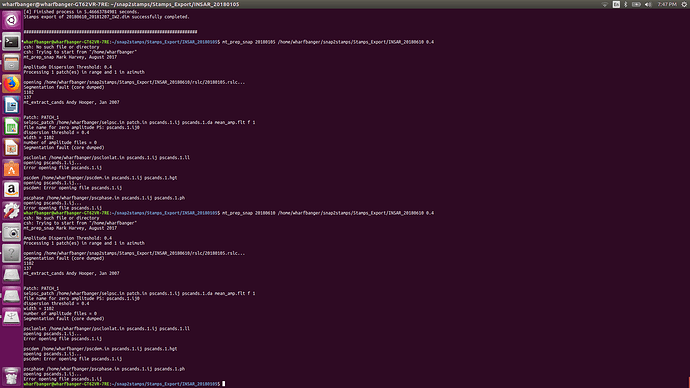
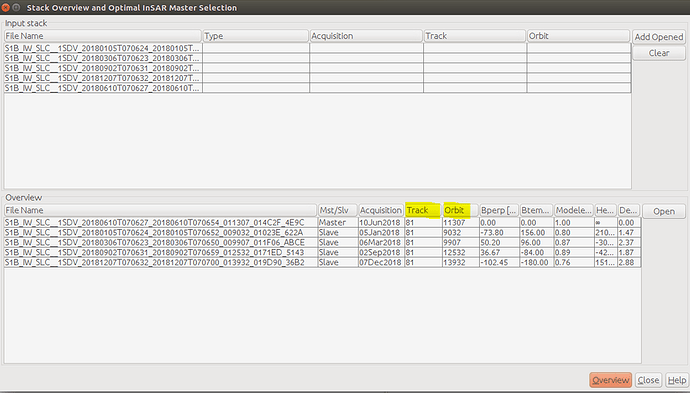
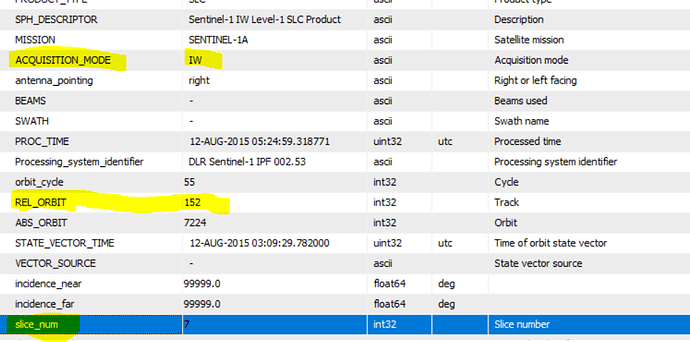
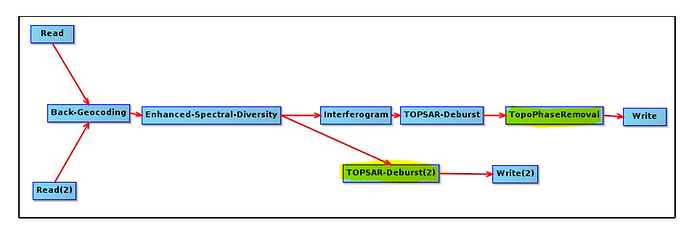
 , it also good to take in the future consideration the effect of LINUX on the processing,
, it also good to take in the future consideration the effect of LINUX on the processing,